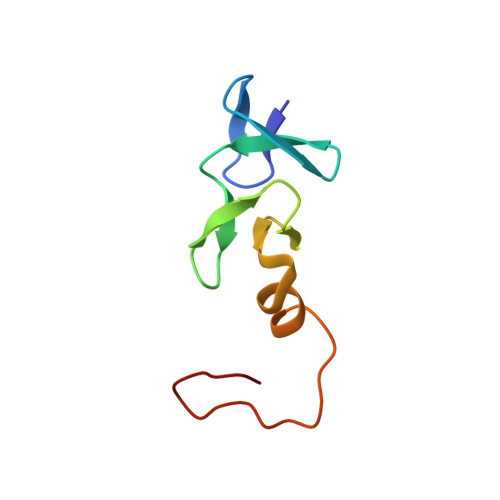
Structure of the cysteine-rich intestinal protein, CRIP.
Perez-Alvarado, G.C., Kosa, J.L., Louis, H.A., Beckerle, M.C., Winge, D.R., Summers, M.F.(1996) J Mol Biology 257: 153-174
- PubMed: 8632452 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0153
- Primary Citation Related Structures:
1IML - PubMed Abstract:
LIM domains are Zn-binding arrays found in a number of proteins involved in the control of cell differentiation, including several developmentally regulated transcription factors and a human proto-oncogene product. The rat cysteine-rich intestinal protein, CRIP, is a 76-residue polypeptide which contains a LIM motif. The solution structure of CRIP has been determined by homonuclear and 1H-15N heteronuclear correlated nuclear magnetic resonance spectroscopy. Structures with individual distance violations of < or = 0.03 angstrom and penalties (squared sum of distance violations) of < or = 0.06 angstrom2 were generated with a total of 500 nuclear Overhauser effect (NOE)-derived distance restraints (averaging 15.6 restraints per refined residue). Superposition of backbone heavy atoms of ordered residues relative to mean atom positions is achieved with pairwise rms deviations of 0.54(+/-0.14) angstrom. As observed previously for a peptide with the sequence of the C-terminal LIM domain from the avian cysteine-rich protein, CRP (cCRP-LIM2), CRIP binds two equivalents of zinc, forming N-terminal CCHC (Cys3, Cys6, His24, Cys27) and C-terminal CCCC (Cys30, Cys33, Cys51, Cys55) modules. The CCHC and CCCC modules in CRIP contain two orthogonally-arrayed antiparallel beta-sheets. The C-terminal end of the CCHC module contains a tight turn and the C terminus of the CCCC module forms an alpha-helix. The modules pack via hydrophobic interactions, forming a compact structure that is similar to that observed for cCRP-LIM2. The most significant differences between the structures occur at the CCHC module-CCCC module interface, which results in a difference in the relative orientations of the modules, and at the C terminus where the alpha-helix appears to be packed more tightly against the preceding antiparallel beta-sheet. The greater abundance of NOE information obtained for CRIP relative to cCRP-LIM2, combined with the analysis of J-coupling and proton chemical shift data, have allowed a more detailed evaluation of the molecular level interactions that stabilize the fold of the LIM motif.
- Howard Hughes Medical Institute, Department of Chemistry and Biochemistry, University of Maryland Baltimore County 21228, USA.
Organizational Affiliation: