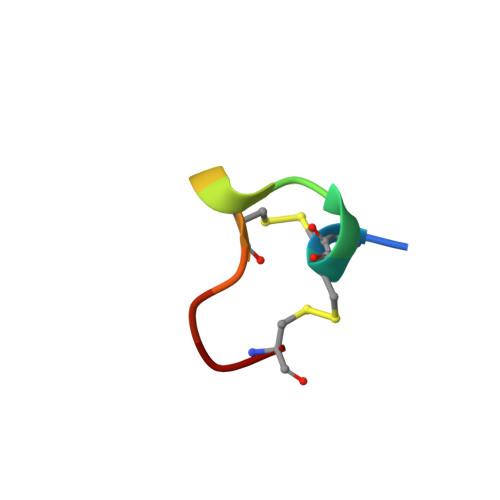
NMR solution structure of alpha-conotoxin ImI and comparison to other conotoxins specific for neuronal nicotinic acetylcholine receptors.
Rogers, J.P., Luginbuhl, P., Shen, G.S., McCabe, R.T., Stevens, R.C., Wemmer, D.E.(1999) Biochemistry 38: 3874-3882
- PubMed: 10194298 Search on PubMed
- DOI: https://doi.org/10.1021/bi9826254
- Primary Citation Related Structures:
1IM1 - PubMed Abstract:
Alpha-Conotoxins, peptides produced by predatory species of Conus marine snails, are potent antagonists of nicotinic acetylcholine receptors (nAChRs), ligand-gated ion channels involved in synaptic transmission. We determined the NMR solution structure of the smallest known alpha-conotoxin, ImI, a 12 amino acid peptide that binds specifically to neuronal alpha7-containing nAChRs in mammals. Calculation of the structure was based on a total of 80 upper distance constraints and 31 dihedral angle constraints resulting in 20 representative conformers with an average pairwise rmsd of 0.44 A from the mean structure for the backbone atoms N, Calpha, and C' of residues 2-11. The structure of ImI is characterized by two compact loops, defined by two disulfide bridges, which form distinct subdomains separated by a deep cleft. Two short 310-helical regions in the first loop are followed by a C-terminal beta-turn in the second. The two disulfide bridges and Ala 9 form a rigid hydrophobic core, orienting the other amino acid side chains toward the surface. Comparison of the three-dimensional structure of ImI to those of the larger, 16 amino acid alpha-conotoxins PnIA, PnIB, MII, and EpI-also specific for neuronal nAChRs-reveals remarkable similarity in local backbone conformations and relative solvent-accessible surface areas. The core scaffold is conserved in all five conotoxins, whereas the residues in solvent-exposed positions are highly variable. The second helical region, and the specific amino acids that the helix exposes to solvent, may be particularly important for binding and selectivity. This comparative analysis provides a three-dimensional structural basis for interpretation of mutagenesis data and structure-activity relationships for ImI as well other neuronal alpha-conotoxins.
- Department of Chemistry, University of California, Berkeley 94720, USA.
Organizational Affiliation: