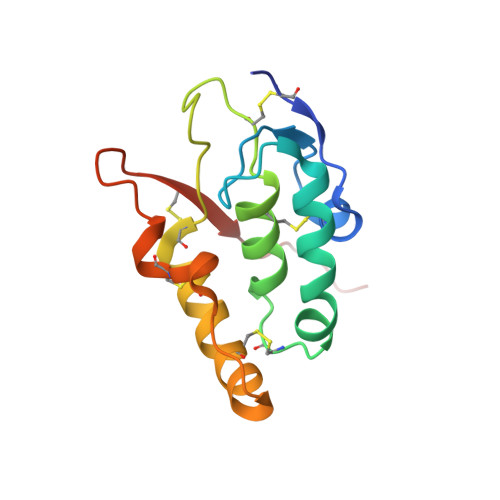
Insights into Wnt binding and signalling from the structures of two Frizzled cysteine-rich domains.
Dann III, C.E., Hsieh, J.C., Rattner, A., Sharma, D., Nathans, J., Leahy, D.J.(2001) Nature 412: 86-90
- PubMed: 11452312 Search on PubMed
- DOI: https://doi.org/10.1038/35083601
- Primary Citation Related Structures:
1IJX, 1IJY - PubMed Abstract:
Members of the Frizzled family of seven-pass transmembrane proteins serve as receptors for Wnt signalling proteins. Wnt proteins have important roles in the differentiation and patterning of diverse tissues during animal development, and inappropriate activation of Wnt signalling pathways is a key feature of many cancers. An extracellular cysteine-rich domain (CRD) at the amino terminus of Frizzled proteins binds Wnt proteins, as do homologous domains in soluble proteins-termed secreted Frizzled-related proteins-that function as antagonists of Wnt signalling. Recently, an LDL-receptor-related protein has been shown to function as a co-receptor for Wnt proteins and to bind to a Frizzled CRD in a Wnt-dependent manner. To investigate the molecular nature of the Wnt signalling complex, we determined the crystal structures of the CRDs from mouse Frizzled 8 and secreted Frizzled-related protein 3. Here we show a previously unknown protein fold, and the design and interpretation of CRD mutations that identify a Wnt-binding site. CRDs exhibit a conserved dimer interface that may be a feature of Wnt signalling. This work provides a framework for studies of homologous CRDs in proteins including muscle-specific kinase and Smoothened, a component of the Hedgehog signalling pathway.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD 21205, USA.
Organizational Affiliation: