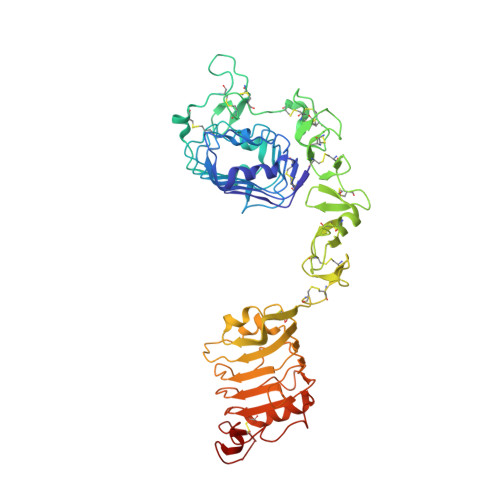
Crystal structure of the first three domains of the type-1 insulin-like growth factor receptor.
Garrett, T.P., McKern, N.M., Lou, M., Frenkel, M.J., Bentley, J.D., Lovrecz, G.O., Elleman, T.C., Cosgrove, L.J., Ward, C.W.(1998) Nature 394: 395-399
- PubMed: 9690478 Search on PubMed
- DOI: https://doi.org/10.1038/28668
- Primary Citation Related Structures:
1IGR - PubMed Abstract:
The type-1 insulin-like growth-factor receptor (IGF-1R) and insulin receptor (IR) are closely related members of the tyrosine-kinase receptor superfamily. IR is essential for glucose homeostasis, whereas IGF-1R is involved in both normal growth and development and malignant transformation. Homologues of these receptors are found in animals as simple as cnidarians. The epidermal growth-factor receptor (EGFR) family is closely related to the IR family and has significant sequence identity to the extracellular portion we describe here. We now present the structure of the first three domains of IGF-IR (L1-Cys-rich-L2) determined to 2.6 A resolution. The L domains each consist of a single-stranded right-handed beta-helix. The Cys-rich region is composed of eight disulphide-bonded modules, seven of which form a rod-shaped domain with modules associated in an unusual manner. The three domains surround a central space of sufficient size to accommodate a ligand molecule. Although the fragment (residues 1-462) does not bind ligand, many of the determinants responsible for hormone binding and ligand specificity map to this central site. This structure therefore shows how the IR subfamily might interact with their ligands.
- Biomolecular Research Institute, Parkville, Victoria, Australia. tom.barrett@bioresi.com.au
Organizational Affiliation: