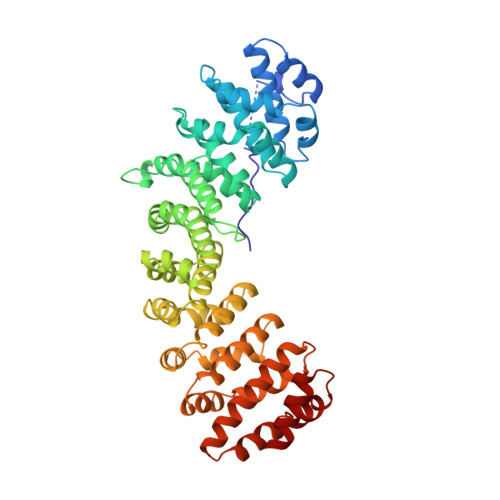
Autoinhibition by an internal nuclear localization signal revealed by the crystal structure of mammalian importin alpha.
Kobe, B.(1999) Nat Struct Biol 6: 388-397
- PubMed: 10201409 Search on PubMed
- DOI: https://doi.org/10.1038/7625
- Primary Citation Related Structures:
1IAL - PubMed Abstract:
Importin alpha is the nuclear import receptor that recognizes classical monopartite and bipartite nuclear localization signals (NLSs). The structure of mouse importin alpha has been determined at 2.5 A resolution. The structure shows a large C-terminal domain containing armadillo repeats, and a less structured N-terminal importin beta-binding domain containing an internal NLS bound to the NLS-binding site. The structure explains the regulatory switch between the cytoplasmic, high-affinity form, and the nuclear, low-affinity form for NLS binding of the nuclear import receptor predicted by the current models of nuclear import. Importin beta conceivably converts the low- to high-affinity form by binding to a site overlapping the autoinhibitory sequence. The structure also has implications for understanding NLS recognition, and the structures of armadillo and HEAT repeats.
- Structural Biology Laboratory, St. Vincent's Institute of Medical Research, Fitzroy, Victoria, Australia. B.Kobe@medicine.unimelb.edu.au
Organizational Affiliation: