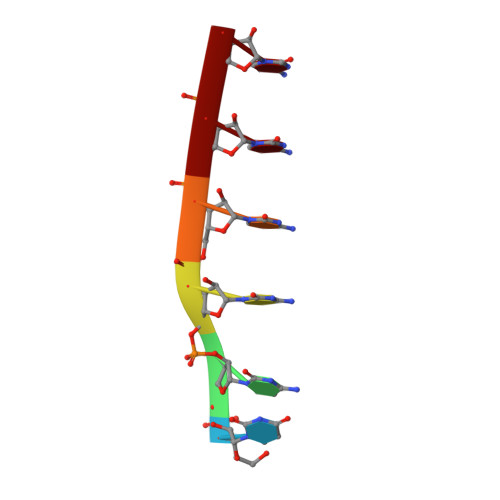
The RNA i-motif.
Snoussi, K., Nonin-Lecomte, S., Leroy, J.L.(2001) J Mol Biology 309: 139-153
- PubMed: 11491284 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4618
- Primary Citation Related Structures:
1I9K - PubMed Abstract:
Oligodeoxynucleotides with stretches of cytidine residues associate into a four-stranded structure, the i-motif, in which two head-to-tail, intercalated, parallel-stranded duplexes are held together by hemiprotonated C.C+ pairs. We have investigated the possibility of forming an i-motif structure with C-rich ribonucleic acids. The four C-rich RNAs studied, r(UC5), r(C5), r(C5U) and r(UC3), associate into multiple intercalated structures at acidic pH. r(UC5) forms two i-motif structures that differ by their intercalation topologies. We report on a structural study of the main form and we analyze the small conformational differences found by comparison with the DNA i-motif. The stacking topology of the main structure avoids one of the six 2'-OH/2'-OH repulsive contacts expected in a fully intercalated structure. The C3'-endo pucker of the RNA sugars and the orientation of the intercalated C.C+ pairs result in a modest widening of the narrow grooves at the steps where the hydroxyl groups are in close contact. The free energy of the RNA i-motif, on average -4 kJ mol(-1) per C.C+ pair, is half of the value found in DNA i-motif structures.
- Groupe de Biophysique de l' Ecole Polytechnique et de l'UMR 7643 du CNRS, Palasieau, France.
Organizational Affiliation: