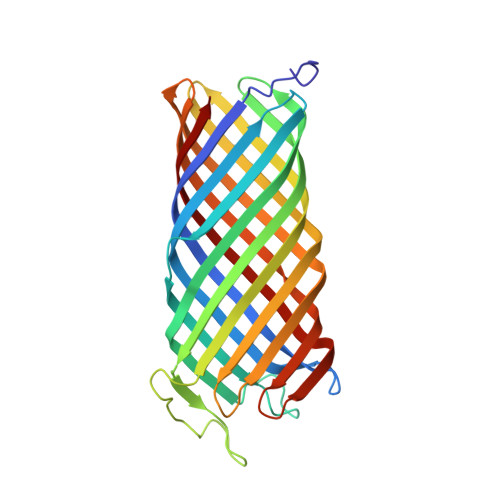
Crystal structure of the outer membrane protease OmpT from Escherichia coli suggests a novel catalytic site.
Vandeputte-Rutten, L., Kramer, R.A., Kroon, J., Dekker, N., Egmond, M.R., Gros, P.(2001) EMBO J 20: 5033-5039
- PubMed: 11566868 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/20.18.5033
- Primary Citation Related Structures:
1I78 - PubMed Abstract:
OmpT from Escherichia coli belongs to a family of highly homologous outer membrane proteases, known as omptins, which are implicated in the virulence of several pathogenic Gram-negative bacteria. Here we present the crystal structure of OmpT, which shows a 10-stranded antiparallel beta-barrel that protrudes far from the lipid bilayer into the extracellular space. We identified a putative binding site for lipopolysaccharide, a molecule that is essential for OmpT activity. The proteolytic site is located in a groove at the extracellular top of the vase-shaped beta-barrel. Based on the constellation of active site residues, we propose a novel proteolytic mechanism, involving a His-Asp dyad and an Asp-Asp couple that activate a putative nucleophilic water molecule. The active site is fully conserved within the omptin family. Therefore, the structure described here provides a sound basis for the design of drugs against omptin-mediated bacterial pathogenesis. Coordinates are in the Protein Data Bank (accession No. 1I78)
- Department of Crystal and Structural Chemistry, Bijvoet Center for Biomolecular Research, Utrecht University, Padualaan 8, 3584 CH Utrecht, The Netherlands.
Organizational Affiliation: