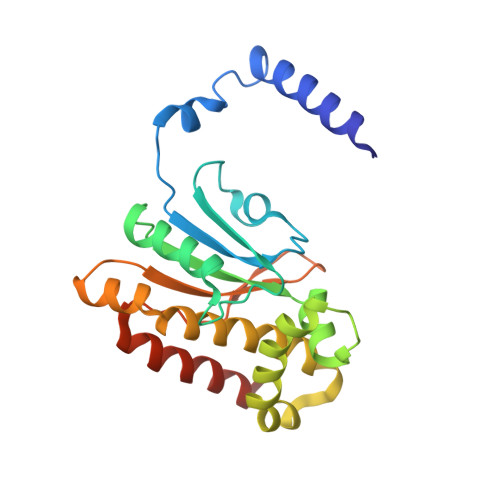
Crystal structure of E. coli beta-carbonic anhydrase, an enzyme with an unusual pH-dependent activity.
Cronk, J.D., Endrizzi, J.A., Cronk, M.R., O'neill, J.W., Zhang, K.Y.(2001) Protein Sci 10: 911-922
- PubMed: 11316870 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.46301
- Primary Citation Related Structures:
1I6O, 1I6P - PubMed Abstract:
Carbonic anhydrases fall into three distinct evolutionary and structural classes: alpha, beta, and gamma. The beta-class carbonic anhydrases (beta-CAs) are widely distributed among higher plants, simple eukaryotes, eubacteria, and archaea. We have determined the crystal structure of ECCA, a beta-CA from Escherichia coli, to a resolution of 2.0 A. In agreement with the structure of the beta-CA from the chloroplast of the red alga Porphyridium purpureum, the active-site zinc in ECCA is tetrahedrally coordinated by the side chains of four conserved residues. These results confirm the observation of a unique pattern of zinc ligation in at least some beta-CAS: The absence of a water molecule in the inner coordination sphere is inconsistent with known mechanisms of CA activity. ECCA activity is highly pH-dependent in the physiological range, and its expression in yeast complements an oxygen-sensitive phenotype displayed by a beta-CA-deletion strain. The structural and biochemical characterizations of ECCA presented here and the comparisons with other beta-CA structures suggest that ECCA can adopt two distinct conformations displaying widely divergent catalytic rates.
- Division of Basic Sciences, Fred Hutchinson Cancer Research Center, Seattle, Washington 98109-1024, USA.
Organizational Affiliation: