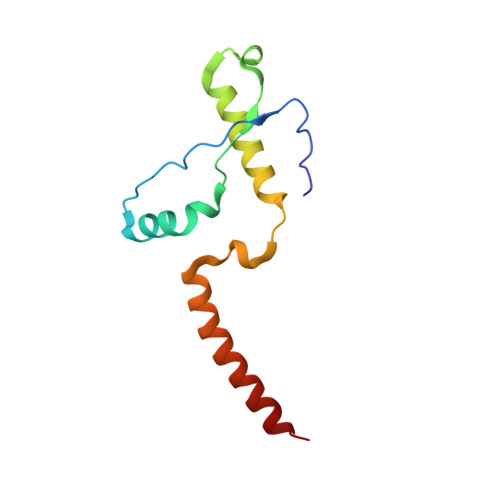
Crystal structure of the human prion protein reveals a mechanism for oligomerization.
Knaus, K.J., Morillas, M., Swietnicki, W., Malone, M., Surewicz, W.K., Yee, V.C.(2001) Nat Struct Biol 8: 770-774
- PubMed: 11524679 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0901-770
- Primary Citation Related Structures:
1I4M - PubMed Abstract:
The pathogenesis of transmissible encephalopathies is associated with the conversion of the cellular prion protein, PrP(C), into a conformationally altered oligomeric form, PrP(Sc). Here we report the crystal structure of the human prion protein in dimer form at 2 A resolution. The dimer results from the three-dimensional swapping of the C-terminal helix 3 and rearrangement of the disulfide bond. An interchain two-stranded antiparallel beta-sheet is formed at the dimer interface by residues that are located in helix 2 in the monomeric NMR structures. Familial prion disease mutations map to the regions directly involved in helix swapping. This crystal structure suggests that oligomerization through 3D domain-swapping may constitute an important step on the pathway of the PrP(C) --> PrP(Sc) conversion.
- Department of Molecular Cardiology and Center for Structural Biology, Lerner Research Institute, Cleveland Clinic Foundation, 9500 Euclid Avenue NB20, Cleveland, Ohio 44195, USA.
Organizational Affiliation: