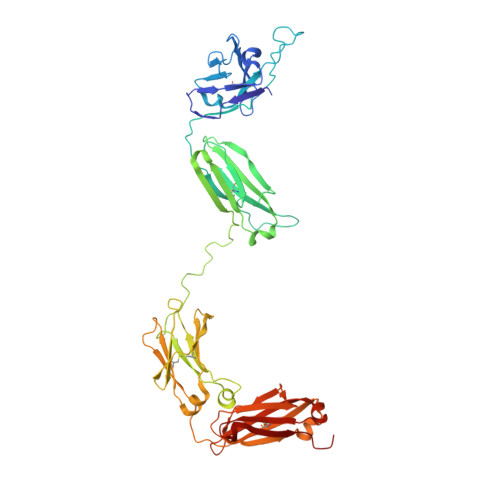
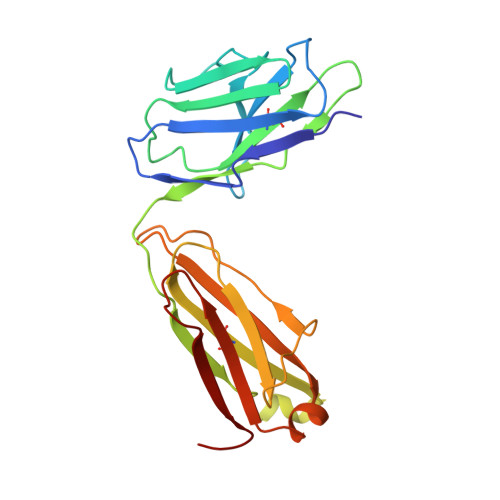
Crystal structure of a neutralizing human IGG against HIV-1: a template for vaccine design.
Saphire, E.O., Parren, P.W., Pantophlet, R., Zwick, M.B., Morris, G.M., Rudd, P.M., Dwek, R.A., Stanfield, R.L., Burton, D.R., Wilson, I.A.(2001) Science 293: 1155-1159
- PubMed: 11498595 Search on PubMed
- DOI: https://doi.org/10.1126/science.1061692
- Primary Citation Related Structures:
1HZH - PubMed Abstract:
We present the crystal structure at 2.7 angstrom resolution of the human antibody IgG1 b12. Antibody b12 recognizes the CD4-binding site of human immunodeficiency virus-1 (HIV-1) gp120 and is one of only two known antibodies against gp120 capable of broad and potent neutralization of primary HIV-1 isolates. A key feature of the antibody-combining site is the protruding, finger-like long CDR H3 that can penetrate the recessed CD4-binding site of gp120. A docking model of b12 and gp120 reveals severe structural constraints that explain the extraordinary challenge in eliciting effective neutralizing antibodies similar to b12. The structure, together with mutagenesis studies, provides a rationale for the extensive cross-reactivity of b12 and a valuable framework for the design of HIV-1 vaccines capable of eliciting b12-like activity.
- Department of Molecular Biology, Department of Immunology, The Skaggs Institute for Chemical Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: