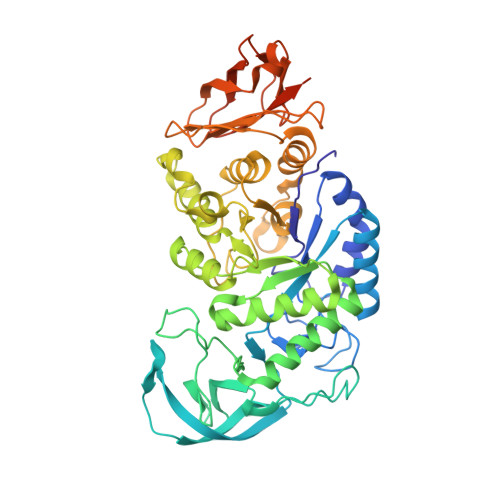
Crystal structure of Bacillus stearothermophilus alpha-amylase: possible factors determining the thermostability.
Suvd, D., Fujimoto, Z., Takase, K., Matsumura, M., Mizuno, H.(2001) J Biochem 129: 461-468
- PubMed: 11226887 Search on PubMed
- DOI: https://doi.org/10.1093/oxfordjournals.jbchem.a002878
- Primary Citation Related Structures:
1HVX - PubMed Abstract:
The crystal structure of a thermostable alpha-amylase from Bacillus stearothermophilus (BSTA) has been determined at 2.0 A resolution. The main-chain fold is almost identical to that of the known crystal structure of Bacillus licheniformis alpha-amylase (BLA). BLA is known to be more stable than BSTA. A structural comparison between the crystal structures of BSTA and BLA showed significant differences that may account for the difference in their thermostabilities, as follows. (i) The two-residue insertion in BSTA, Ile181-Gly182, pushes away the spatially contacting region including Asp207, which corresponds to Ca(2+)-coordinating Asp204 in BLA. As a result, Asp207 cannot coordinate the Ca(2+). (ii) BSTA contains nine fewer hydrogen bonds than BLA, which costs about 12 kcal/mol. This tendency is prominent in the (beta/alpha)(8)-barrel, where 10 fewer hydrogen bonds were observed in BSTA. BLA forms a denser hydrogen bond network in the inter-helical region, which may stabilize alpha-helices in the barrel. (iii) A few small voids observed in the alpha-helical region of the (beta/alpha)(8)-barrel in BSTA decrease inter-helical compactness and hydrophobic interactions. (iv) The solvent-accessible surface area of charged residues in BLA is about two times larger than that in BSTA.
- Institute of Applied Biochemistry, University of Tsukuba, Tsukuba, Ibaraki 305-8572, Japan. mizuno@abr.affrc.go.jp
Organizational Affiliation: