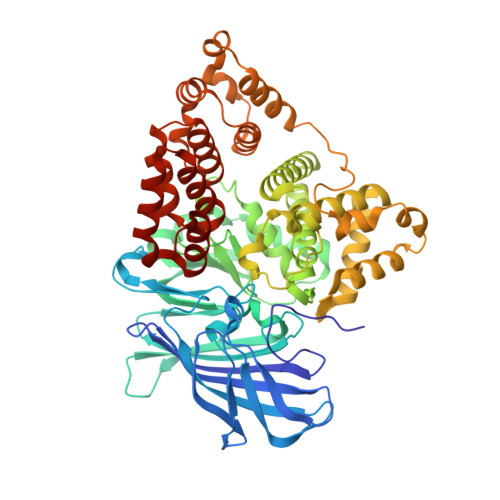
Crystal structure of human leukotriene A(4) hydrolase, a bifunctional enzyme in inflammation.
Thunnissen, M.M., Nordlund, P., Haeggstrom, J.Z.(2001) Nat Struct Biol 8: 131-135
- PubMed: 11175901 Search on PubMed
- DOI: https://doi.org/10.1038/84117
- Primary Citation Related Structures:
1HS6 - PubMed Abstract:
Leukotriene (LT) A(4) hydrolase/aminopeptidase (LTA4H) is a bifunctional zinc enzyme that catalyzes the biosynthesis of LTB4, a potent lipid chemoattractant involved in inflammation, immune responses, host defense against infection, and PAF-induced shock. The high resolution crystal structure of LTA4H in complex with the competitive inhibitor bestatin reveals a protein folded into three domains that together create a deep cleft harboring the catalytic Zn(2+) site. A bent and narrow pocket, shaped to accommodate the substrate LTA(4), constitutes a highly confined binding region that can be targeted in the design of specific anti-inflammatory agents. Moreover, the structure of the catalytic domain is very similar to that of thermolysin and provides detailed insight into mechanisms of catalysis, in particular the chemical strategy for the unique epoxide hydrolase reaction that generates LTB(4).
- Department of Biochemistry, University of Stockholm, Arrhenius Laboratories A4, S-106 91 Stockholm, Sweden. marjo@dbb.su.se
Organizational Affiliation: