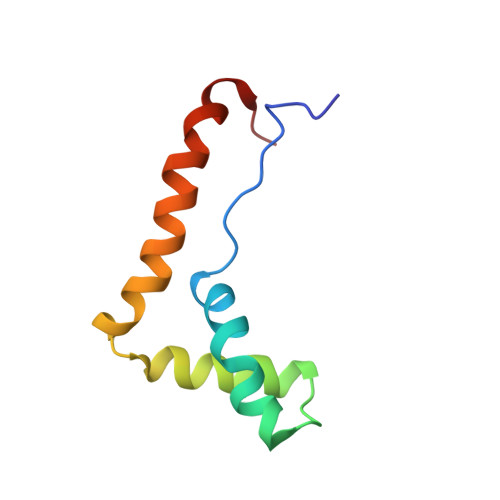
Structure of the HMG box motif in the B-domain of HMG1.
Weir, H.M., Kraulis, P.J., Hill, C.S., Raine, A.R., Laue, E.D., Thomas, J.O.(1993) EMBO J 12: 1311-1319
- PubMed: 8467791 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/j.1460-2075.1993.tb05776.x
- Primary Citation Related Structures:
1HME, 1HMF - PubMed Abstract:
The conserved, abundant chromosomal protein HMG1 consists of two highly homologous, folded, basic DNA-binding domains, each of approximately 80 amino acid residues, and an acidic C-terminal tail. Each folded domain represents an 'HMG box', a sequence motif recently recognized in certain sequence-specific DNA-binding proteins and which also occurs in abundant HMG1-like proteins that bind to DNA without sequence specificity. The HMG box is defined by a set of highly conserved residues (most distinctively aromatic and basic) and appears to define a novel DNA-binding structural motif. We have expressed the HMG box region of the B-domain of rat HMG1 (residues 88-164 of the intact protein) in Escherichia coli and we describe here the determination of its structure by 2D 1H-NMR spectroscopy. There are three alpha-helices (residues 13-29, 34-48 and 50-74), which together account for approximately 75% of the total residues and contain many of the conserved basic and aromatic residues. Strikingly, the molecule is L-shaped, the angle of approximately 80 degrees between the two arms being defined by a cluster of conserved, predominantly aromatic, residues. The distinctive shape of the HMG box motif, which is distinct from hitherto characterized DNA-binding motifs, may be significant in relation to its recognition of four-way DNA junctions.
- Department of Biochemistry, University of Cambridge, UK.
Organizational Affiliation: