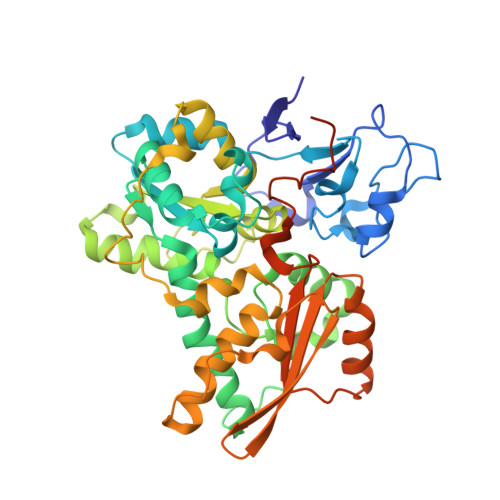
Desulfovibrio desulfuricans iron hydrogenase: the structure shows unusual coordination to an active site Fe binuclear center.
Nicolet, Y., Piras, C., Legrand, P., Hatchikian, C.E., Fontecilla-Camps, J.C.(1999) Structure 7: 13-23
- PubMed: 10368269 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(99)80005-7
- Primary Citation Related Structures:
1HFE - PubMed Abstract:
Many microorganisms have the ability to either oxidize molecular hydrogen to generate reducing power or to produce hydrogen in order to remove low-potential electrons. These reactions are catalyzed by two unrelated enzymes: the Ni-Fe hydrogenases and the Fe-only hydrogenases. We report here the structure of the heterodimeric Fe-only hydrogenase from Desulfovibrio desulfuricans - the first for this class of enzymes. With the exception of a ferredoxin-like domain, the structure represents a novel protein fold. The so-called H cluster of the enzyme is composed of a typical [4Fe-4S] cubane bridged to a binuclear active site Fe center containing putative CO and CN ligands and one bridging 1, 3-propanedithiol molecule. The conformation of the subunits can be explained by the evolutionary changes that have transformed monomeric cytoplasmic enzymes into dimeric periplasmic enzymes. Plausible electron- and proton-transfer pathways and a putative channel for the access of hydrogen to the active site have been identified. The unrelated active sites of Ni-Fe and Fe-only hydrogenases have several common features: coordination of diatomic ligands to an Fe ion; a vacant coordination site on one of the metal ions representing a possible substrate-binding site; a thiolate-bridged binuclear center; and plausible proton- and electron-transfer pathways and substrate channels. The diatomic coordination to Fe ions makes them low spin and favors low redox states, which may be required for catalysis. Complex electron paramagnetic resonance signals typical of Fe-only hydrogenases arise from magnetic interactions between the [4Fe-4S] cluster and the active site binuclear center. The paucity of protein ligands to this center suggests that it was imported from the inorganic world as an already functional unit.
- Laboratoire de Cristallographie et de Cristallogènese des Protéines, Institut de Biologie Structurale Jean-Pierre Ebel, CEA-CNRS 41, Grenoble, France.
Organizational Affiliation: