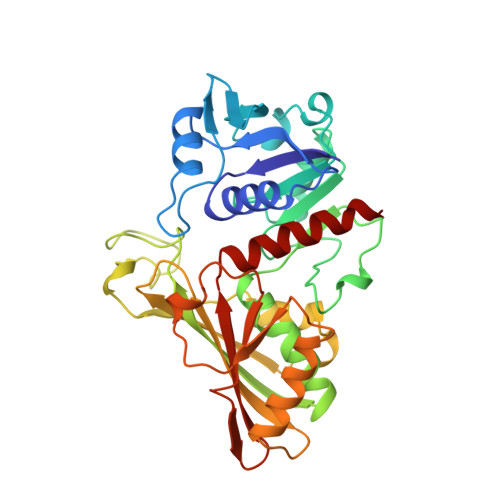
The crystal structure of holo-glyceraldehyde-3-phosphate dehydrogenase from the hyperthermophilic bacterium Thermotoga maritima at 2.5 A resolution.
Korndorfer, I., Steipe, B., Huber, R., Tomschy, A., Jaenicke, R.(1995) J Mol Biology 246: 511-521
- PubMed: 7877172 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1994.0103
- Primary Citation Related Structures:
1HDG - PubMed Abstract:
The crystal structure of holo-glyceraldehyde-3-phosphate dehydrogenase from the hyperthermophile Thermotoga maritima was determined by Patterson search methods using the known structure of the Bacillus stearothermophilus enzyme. The structure was refined at a resolution of 2.5 A to an R-factor of 16.63% for 26289 reflections between 8.0 A an 2.5 A with F > 2 sigma(F). The crystallographic asymmetric unit contains two monomers related by approximate 2-fold symmetry and a tetramer is built up by crystallographic symmetry. The root-mean-square deviation of Ca positions of glyceraldehyde-3-phosphate dehydrogenase from T. maritima and B. stearothermophilus is 0.83 A in the NAD+ binding domains and smaller close to the cofactor. In contrast, the largest deviations in the catalytic domains are found at residues involved in coordination of sulphate ion SO4 339, which most likely marks the site of the attacking inorganic phosphate ion in catalysis. A large number of extra salt-bridges may be an important factor contributing to the high thermostability of this protein.
- Max-Planck Institut für Biochemie, Martinsried, FRG.
Organizational Affiliation: