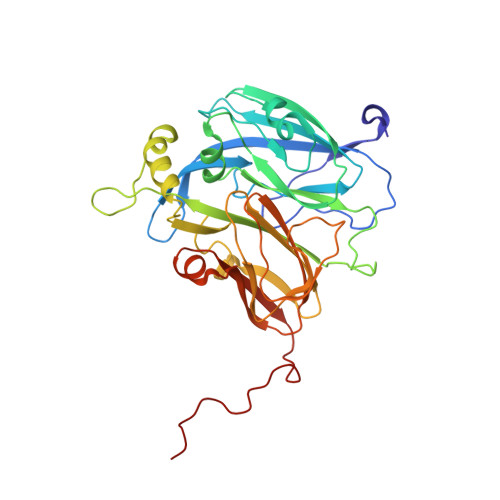
X-Ray Structure of a Blue Copper Nitrite Reductase at High Ph and in Copper-Free Form at 1.9 A Resolution
Ellis, M.J., Dodd, F.E., Strange, R.W., Prudencio, M., Sawers, G., Eady, R.R., Hasnain, S.S.(2001) Acta Crystallogr D Biol Crystallogr 57: 1110
- PubMed: 11468394 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444901008654
- Primary Citation Related Structures:
1HAU, 1HAW - PubMed Abstract:
Copper-containing nitrite reductases possess a trimeric structure where the catalytic Cu site, located at the monomer-monomer interface, resembles the catalytic sites of a number of Zn enzymes. Nitrite reductase from Alcaligenes xylosoxidans has optimum activity at pH 5.2 which decreases to a negligible level at pH 8. The structure of this nitrite reductase has previously been determined at pH 4.6. It has now been crystallized under new conditions at pH 8.5. Its crystallographic structure provides a structural explanation for the greatly reduced activity of the enzyme at high pH. Characterization of overexpressed protein in solution by EXAFS suggested that the protein lacked Cu in the catalytic type 2 Cu site and that the site was most probably occupied by Zn. Using the anomalous signals from Cu and Zn, the crystal structure revealed that the expressed protein was devoid of Cu in the catalytic site and that only a trace amount (<10%) of Zn was present at this site in the crystal. Despite the close structural similarity of the catalytic site to a number of Zn enzymes, these data suggest that Zn, if it binds at the catalytic copper site, binds weakly in nitrite reductase.
- Synchrotron Radiation Department, CLRC Daresbury Laboratory, Warrington WA4 4AD, England.
Organizational Affiliation: