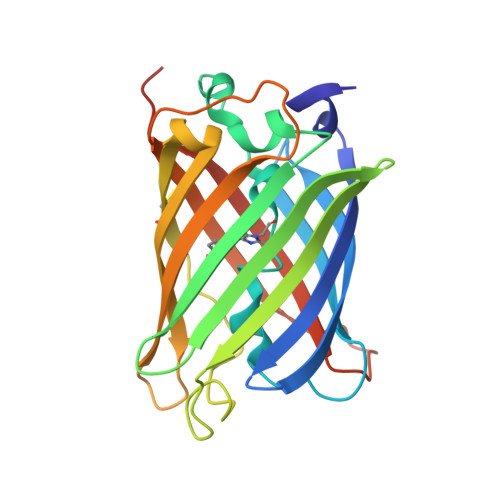
Shedding Light on Disulfide Bond Formation: Engineering a Redox Switch in Green Fluorescent Protein
Ostergaard, H., Henriksen, A., Hansen, F.G., Winther, J.R.(2001) EMBO J 20: 5853
- PubMed: 11689426 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/20.21.5853
- Primary Citation Related Structures:
1H6R - PubMed Abstract:
To visualize the formation of disulfide bonds in living cells, a pair of redox-active cysteines was introduced into the yellow fluorescent variant of green fluorescent protein. Formation of a disulfide bond between the two cysteines was fully reversible and resulted in a >2-fold decrease in the intrinsic fluorescence. Inter conversion between the two redox states could thus be followed in vitro as well as in vivo by non-invasive fluorimetric measurements. The 1.5 A crystal structure of the oxidized protein revealed a disulfide bond-induced distortion of the beta-barrel, as well as a structural reorganization of residues in the immediate chromophore environment. By combining this information with spectroscopic data, we propose a detailed mechanism accounting for the observed redox state-dependent fluorescence. The redox potential of the cysteine couple was found to be within the physiological range for redox-active cysteines. In the cytoplasm of Escherichia coli, the protein was a sensitive probe for the redox changes that occur upon disruption of the thioredoxin reductive pathway.
- Section of Molecular Microbiology, BioCentrum-DTU, Technical University of Denmark, Building 301, DK-2800 Lyngby, Denmark.
Organizational Affiliation: