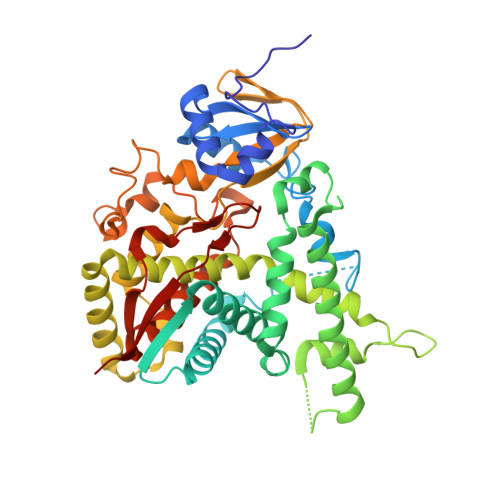
Estriol Bound and Ligand-Free Structures of Sterol 14Alpha-Demethylase.
Podust, L.M., Yermalitskaya, L.V., Lepesheva, G.I., Podust, V.N., Dalmasso, E.A., Waterman, M.R.(2004) Structure 12: 1937
- PubMed: 15530358 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.08.009
- Primary Citation Related Structures:
1H5Z, 1X8V - PubMed Abstract:
Sterol 14alpha-demethylases (CYP51) are essential enzymes in sterol biosynthesis in eukaryotes and drug targets in antifungal therapy. Here, we report CYP51 structures in ligand-free and estriol bound forms. Using estriol as a probe, we determined orientation of the substrate in the active site, elucidated protein contacts with the invariant 3beta-hydroxy group of a sterol, and identified F78 as a key discriminator between 4alpha-methylated and 4alpha,beta-dimethylated substrates. Analysis of CYP51 dynamics revealed that the C helix undergoes helix-coil transition upon binding and dissociation of a ligand. Loss of helical structure of the C helix in the ligand-free form results in an unprecedented opening of the substrate binding site. Upon binding of estriol, the BC loop loses contacts with molecular surface and tends to adopt a closed conformation. A mechanism for azole resistance in the yeast pathogen Candida albicans associated with mutations in the ERG11 gene encoding CYP51 is suggested based on CYP51 protein dynamics.
- Department of Biochemistry, Vanderbilt University School of Medicine, Nashville, TN 37232, USA. larissa.m.podust@vanderbilt.edu
Organizational Affiliation: