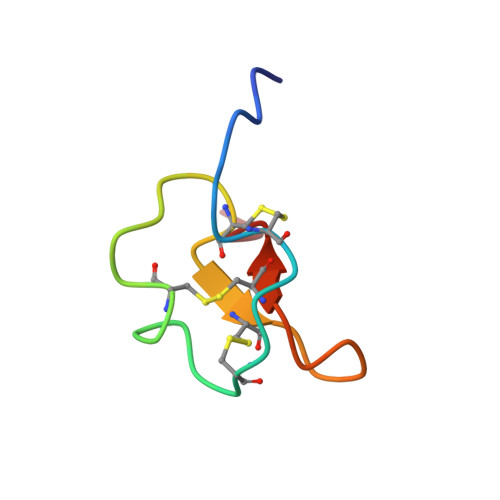
Structure and Dynamics of the Potato Carboxypeptidase Inhibitor by 1H and 15N NMR.
Gonzalez, C., Neira, J.L., Ventura, S., Bronsoms, S., Rico, M., Aviles, F.X.(2003) Proteins 50: 410
- PubMed: 12557184 Search on PubMed
- DOI: https://doi.org/10.1002/prot.10291
- Primary Citation Related Structures:
1H20 - PubMed Abstract:
The solution structure and backbone dynamics of the recombinant potato carboxypeptidase inhibitor (PCI) have been characterized by NMR spectroscopy. The structure, determined on the basis of 497 NOE-derived distance constraints, is much better defined than the one reported in a previous NMR study, with an average pairwise backbone root-mean-square deviation of 0.5 A for the well-defined region of the protein, residues 7-37. Many of the side-chains show now well-defined conformations, both in the hydrophobic core and on the surface of the protein. Overall, the solution structure of free PCI is similar to the one that it shows in the crystal of the complex with carboxypeptidase A. However, some local differences are observed in regions 15-21 and 27-29. In solution, the six N-terminal and the two C-terminal residues are rather flexible, as shown by 15N backbone relaxation measurements. The flexibility of the latter segment may have implications in the binding of the inhibitor by the enzyme. All the remaining residues in the protein are essentially rigid (S2 > 0.8) with the exception of two of them at the end of a short 3/10 helix. Despite the small size of the protein, a number of amide protons are protected from exchange with solvent deuterons. The slowest exchanging protons are those in a small two-strand beta-sheet. The unfolding free energies, as calculated from the exchange rates of these protons, are around 5 kcal/mol. Other protected amide protons are located in the segment 7-12, adjacent to the beta-sheet. Although these residues are not in an extended conformation in PCI, the equivalent residues in structurally homologous proteins form a third strand of the central beta-sheet. The amide protons in the 3/10 helix are only marginally protected, indicating that they exchange by a local unfolding mechanism, which is consistent with the increase in flexibility shown by some of its residues. Backbone alignment-based programs for folding recognition, as opposite to disulfide-bond alignments, reveal new proteins of unrelated sequence and function with a similar structure.
- Instituto de Química-Física Rocasolano (C.S.I.C.), Serrano, 119, Madrid, Spain.
Organizational Affiliation: