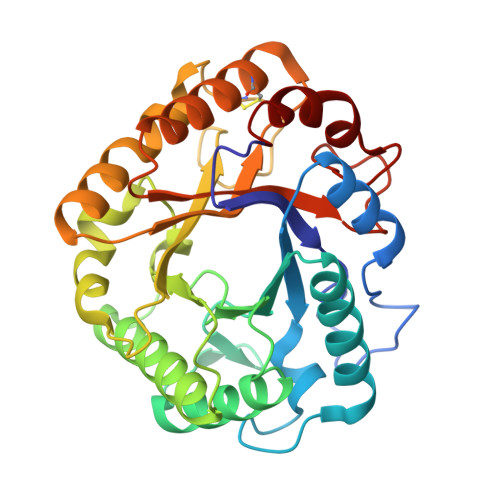
Atomic Resolution Structure of the Major Endoglucanase from Thermoascus Aurantiacus
Van Petegem, F., Vandenberghe, I., Bhat, M.K., Van Beeumen, J.(2002) Biochem Biophys Res Commun 296: 161
- PubMed: 12147244 Search on PubMed
- DOI: https://doi.org/10.1016/s0006-291x(02)00775-1
- Primary Citation Related Structures:
1H1N - PubMed Abstract:
The crystal structure of the major endoglucanase from the thermophilic fungus Thermoascus aurantiacus was determined by single isomorphous replacement at 1.12A resolution. The full sequence supports the classification of the protein in a subgroup of glycoside hydrolase family 5 for which no structural data are available yet. The active site shows eight critical residues, strictly conserved within family 5. In addition, aromatic residues that line the substrate-binding cleft and that are possibly involved in substrate-binding are identified. A number of residues seem to be conserved among members of the subtype, including a disulphide bridge between Cys212 and Cys249.
- Laboratorium voor Eiwitbiochemie en Eiwitengineering, Universiteit Gent, B-9000, Gent, Belgium.
Organizational Affiliation: