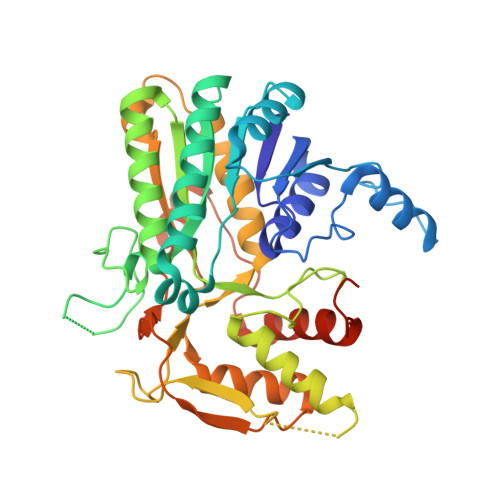
High-Resolution Crystal Structure of Trypanosoma Brucei Udp-Galactose 4'-Epimerase: A Potential Target for Structure-Based Development of Novel Trypanocides
Shaw, M.P., Bond, C.S., Roper, J.R., Gourley, D.G., Ferguson, M.A.J., Hunter, W.N.(2003) Mol Biochem Parasitol 126: 173
- PubMed: 12615316 Search on PubMed
- DOI: https://doi.org/10.1016/s0166-6851(02)00243-8
- Primary Citation Related Structures:
1GY8 - PubMed Abstract:
The crystal structure of UDP-galactose 4'-epimerase from the protozoan parasite Trypanosoma brucei in complex with the cofactor NAD(+) and a fragment of the substrates, UDP, has been determined at 2.0 A resolution (1 A = 0.1 nm). This enzyme, recently proven to be essential for this pathogenic parasite, shares 33% sequence identity with the corresponding enzyme in the human host. Structural comparisons indicate that many of the protein-ligand interactions are conserved between the two enzymes. However, in the UDP-binding pocket there is a non-conservative substitution from Gly237 in the human enzyme to Cys266 in the T. brucei enzyme. Such a significant difference could be exploited by the structure-based design of selective inhibitors using the structure of the trypanosomatid enzyme as a template.
- Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee, Dundee DD1 5EH, UK.
Organizational Affiliation: