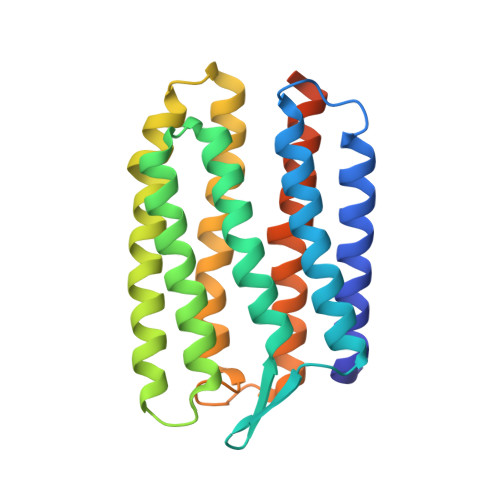
Early Structural Rearrangements in the Photocycle of an Integral Membrane Sensory Receptor
Edman, K., Royant, A., Nollert, P., Maxwell, C.A., Pebay-Peyroula, E., Navarro, J., Neutze, R., Landau, E.M.(2002) Structure 10: 473
- PubMed: 11937052 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00736-0
- Primary Citation Related Structures:
1GU8, 1GUE - PubMed Abstract:
Sensory rhodopsins are the primary receptors of vision in animals and phototaxis in microorganisms. Light triggers the rapid isomerization of a buried retinal chromophore, which the protein both accommodates and amplifies into the larger structural rearrangements required for signaling. We trapped an early intermediate of the photocycle of sensory rhodopsin II from Natronobacterium pharaonis (pSRII) in 3D crystals and determined its X-ray structure to 2.3 A resolution. The observed structural rearrangements were localized near the retinal chromophore, with a key water molecule becoming disordered and the retinal's beta-ionone ring undergoing a prominent movement. Comparison with the early structural rearrangements of bacteriorhodopsin illustrates how modifications in the retinal binding pocket of pSRII allow subtle differences in the early relaxation of photoisomerized retinal.
- Department of Molecular Biotechnology, Chalmers University of Technology, Box 462, S-40530 Gothenburg, Sweden.
Organizational Affiliation: