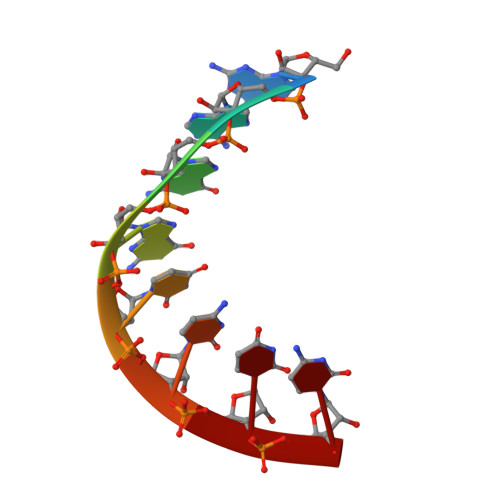
Investigation of the structural basis for thermodynamic stabilities of tandem GU mismatches: solution structure of (rGAGGUCUC)2 by two-dimensional NMR and simulated annealing.
McDowell, J.A., Turner, D.H.(1996) Biochemistry 35: 14077-14089
- PubMed: 8916893 Search on PubMed
- DOI: https://doi.org/10.1021/bi9615710
- Primary Citation Related Structures:
1GUC - PubMed Abstract:
The duplex (rGAGGUCUC)2 contains the motif [sequence: see text] which is unusually stable compared with other symmetric tandem GU mismatches and occurs in the P5 helix of the group I intron of Tetrahymena thermophila. The three-dimensional solution structure of (rGAGGUCUC)2 was determined using two-dimensional NMR and a simulated annealing protocol. The structure is remarkably similar to the A-DNA crystal structure of (dGGGGTCCC)2 [Kneale, G., Brown, T., & Kennard, O. (1985) J. Mol. Biol. 186, 805-814] which contains the analogous motif [sequence: see text]. Incorporation of the [sequence: see text] motif has little effect on backbone torsion angles and helical parameters compared with standard A-form. The only significant departure from A-form is a slight overtwisting 5' of the G in the GU mismatch and a displacement of the mismatches toward the minor groove. Inspection of stacking patterns of this structure and comparison with symmetric tandem GT mismatches in A-DNA oligonucleotides from crystal structure data suggest that electrostatics are important in determining motif stability.
- Department of Chemistry, University of Rochester, New York 14627-0216, USA.
Organizational Affiliation: