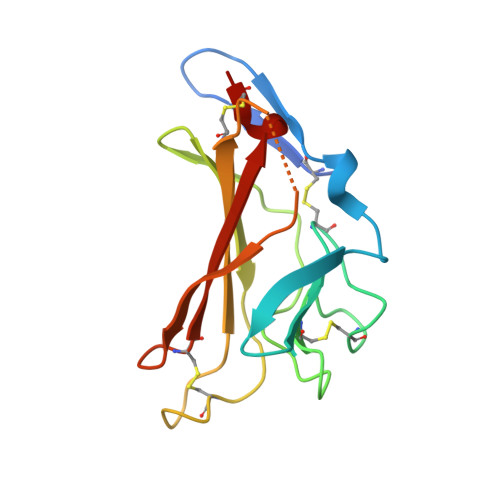
Structure of a Functional Igf2R Fragment Determined from the Anomalous Scattering of Sulfur
Brown, J., Esnouf, R.M., Jones, M.A., Linnell, J., Harlos, K., Hassan, A.B., Jones, E.Y.(2002) EMBO J 21: 1054
- PubMed: 11867533 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/21.5.1054
- Primary Citation Related Structures:
1GP0, 1GP3 - PubMed Abstract:
Insulin-like growth factor II receptor (IGF2R) is a multifunctional cell surface receptor implicated in tumour suppression. Its growth inhibitory activity has been associated with an ability to bind IGF-II. IGF2R contains 15 homologous extracellular domains, with domain 11 primarily responsible for IGF-II binding. We report a 1.4 A resolution crystal structure of domain 11, solved using the anomalous scattering signal of sulfur. The structure consists of two crossed beta-sheets forming a flattened beta-barrel. Structural analysis identifies the putative IGF-II binding site at one end of the beta-barrel whilst crystal lattice contacts suggest a model for the full-length IGF2R extracellular region. The structure factors and coordinates of IGF2R domain 11 have been deposited in the Protein Data Bank (accession codes 1GP0 and 1GP3).
- Cancer Research UK Receptor Structure Research Group, Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Roosevelt Drive, Headington, Oxford OX3 7BN, UK.
Organizational Affiliation: