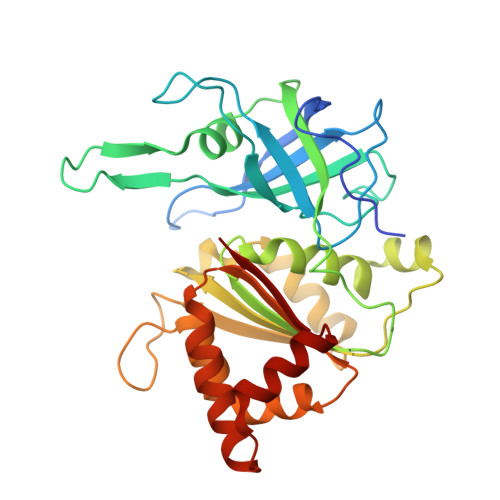
Mechanism of Coenzyme Recognition and Binding Revealed by Crystal Structure Analysis of Ferredoxin-Nadp(+) Reductase Complexed with Nadp(+)
Hermoso, J., Mayoral, T., Faro, M., Gomez-Moreno, C., Sanz-Aparicio, J., Medina, M.(2002) J Mol Biology 319: 1133
- PubMed: 12079352 Search on PubMed
- DOI: https://doi.org/10.1016/S0022-2836(02)00388-1
- Primary Citation Related Structures:
1GJR - PubMed Abstract:
The flavoenzyme ferredoxin-NADP+ reductase (FNR) catalyses the production of NADPH in photosynthesis. The three-dimensional structure of FNR presents two distinct domains, one for binding of the FAD prosthetic group and the other for NADP+ binding. In spite of extensive experiments and different crystallographic approaches, many aspects about how the NADP+ substrate binds to FNR and how the hydride ion is transferred from FAD to NADP+ remain unclear. The structure of an FNR:NADP+ complex from Anabaena has been determined by X-ray diffraction analysis of the cocrystallised units to 2.1 A resolution. Structural perturbation of FNR induced by complex formation produces a narrower cavity in which the 2'-phospho-AMP and pyrophosphate portions of the NADP+ are perfectly bound. In addition, the nicotinamide mononucleotide moiety is placed in a new pocket created near the FAD cofactor with the ribose being in a tight conformation. The crystal structure of this FNR:NADP+ complex obtained by cocrystallisation displays NADP+ in an unusual conformation and can be considered as an intermediate state in the process of coenzyme recognition and binding. Structural analysis and comparison with previously reported complexes allow us to postulate a mechanism which would permit efficient hydride transfer to occur. Besides, this structure gives new insights into the postulated formation of the ferredoxin:FNR:NADP+ ternary complex by prediction of new intermolecular interactions, which could only exist after FNR:NADP+ complex formation. Finally, structural comparison with the members of the broad FNR structural family also provides an explanation for the high specificity exhibited by FNR for NADP+/H versus NAD+/H.
- Grupo de Cristalografía Macromolecular y Biología Estructural, Instituto Química-Física Rocasolano, C.S.I.C., Serrano 119, 28006 Madrid, Spain. xjuan@iqfr.csic.es
Organizational Affiliation: