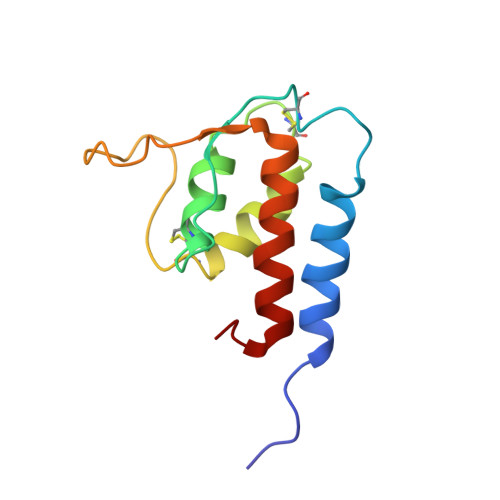
Solution structure of interleukin-13 and insights into receptor engagement
Eisenmesser, E.Z., Horita, D.A., Altieri, A.S., Byrd, R.A.(2001) J Mol Biology 310: 231-241
- PubMed: 11419949 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4765
- Primary Citation Related Structures:
1GA3 - PubMed Abstract:
The complex and interrelated function of the interleukin cytokines relies on a range of pro-inflammatory and anti-inflammatory immune responses mediated by an array of receptors, and there is considerable cross-reactivity for related cytokines. Recent findings continue to elucidate the expression patterns of interleukin receptors associated with a range of diseases, including cancer. We report here the first experimentally determined high-resolution structure of human interleukin-13 (IL-13). The experimental structure is significantly different from an earlier homology model, which could have led to improper estimation of receptor interaction surfaces and design of mutational experiments. Similarities between the presented IL-13 structure and the homologous interleukin-4 (IL-4) are discussed. Additionally, mutation data for IL-4 and IL-13 are analyzed and combined with a detailed structural analysis of the IL-4/IL4Ralpha interface that leads us to postulate interactions at the IL-13/receptor interface. The structural comparison is used to interpret the different affinities for various receptors and establishes the basis for further mutational experiments and antagonist design.
- Macromolecular NMR Section Structural Biophysics Laboratory, National Cancer Institute-Frederick, Frederick, MD 21702-1201, USA.
Organizational Affiliation: