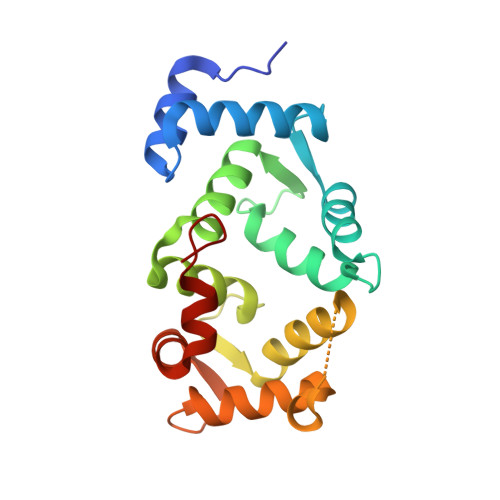
Immunocytochemical localization and crystal structure of human frequenin (neuronal calcium sensor 1).
Bourne, Y., Dannenberg, J., Pollmann, V., Marchot, P., Pongs, O.(2001) J Biological Chem 276: 11949-11955
- PubMed: 11092894 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M009373200
- Primary Citation Related Structures:
1G8I - PubMed Abstract:
Frequenin, a member of a large family of myristoyl-switch calcium-binding proteins, functions as a calcium-ion sensor to modulate synaptic activity and secretion. We show that human frequenin colocalizes with ARF1 GTPase in COS-7 cells and occurs in similar cellular compartments as the phosphatidylinositol-4-OH kinase PI4Kbeta, the mammalian homolog of the yeast kinase PIK1. In addition, the crystal structure of unmyristoylated, calcium-bound human frequenin has been determined and refined to 1.9 A resolution. The overall fold of frequenin resembles those of neurocalcin and the photoreceptor, recoverin, of the same family, with two pairs of calcium-binding EF hands and three bound calcium ions. Despite the similarities, however, frequenin displays significant structural differences. A large conformational shift of the C-terminal region creates a wide hydrophobic crevice at the surface of frequenin. This crevice, which is unique to frequenin and distinct from the myristoyl-binding box of recoverin, may accommodate a yet unknown protein ligand.
- Architecture et Fonction des Macromolécules Biologiques, CNRS UMR-6098, 13402, Marseille Cedex 20, France. yves@afmb.cnrs-mrs.fr
Organizational Affiliation: