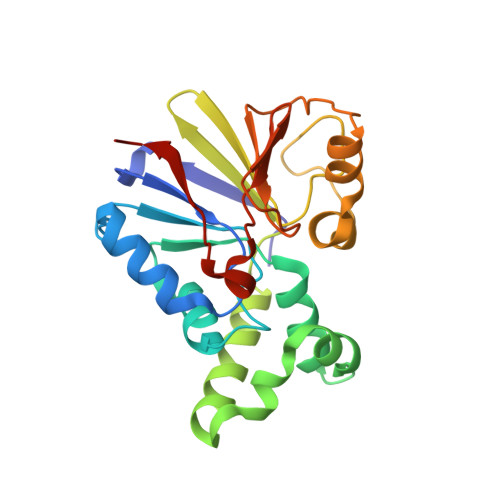
Structure of the bacteriophage lambda Ser/Thr protein phosphatase with sulfate ion bound in two coordination modes.
Voegtli, W.C., White, D.J., Reiter, N.J., Rusnak, F., Rosenzweig, A.C.(2000) Biochemistry 39: 15365-15374
- PubMed: 11112522 Search on PubMed
- DOI: https://doi.org/10.1021/bi0021030
- Primary Citation Related Structures:
1G5B - PubMed Abstract:
The protein phosphatase encoded by bacteriophage lambda (lambda PP) belongs to a family of Ser/Thr phosphatases (Ser/Thr PPases) that includes the eukaryotic protein phosphatases 1 (PP1), 2A (PP2A), and 2B (calcineurin). These Ser/Thr PPases and the related purple acid phosphatases (PAPs) contain a conserved phosphoesterase sequence motif that binds a dinuclear metal center. The mechanisms of phosphoester hydrolysis by these enzymes are beginning to be unraveled. To utilize lambda PP more effectively as a model for probing the catalytic mechanism of the Ser/Thr PPases, we have determined its crystal structure to 2.15 A resolution. The overall fold resembles that of PP1 and calcineurin, including a conserved beta alpha beta alpha beta structure that comprises the phosphoesterase motif. Substrates and inhibitors probably bind in a narrow surface groove that houses the active site dinuclear Mn(II) center. The arrangement of metal ligands is similar to that in PP1, calcineurin, and PAP, and a bound sulfate ion is present in two novel coordination modes. In two of the three molecules in the crystallographic asymmetric unit, sulfate is coordinated to Mn2 in a monodentate, terminal fashion, and the two Mn(II) ions are bridged by a solvent molecule. Two additional solvent molecules are coordinated to Mn1. In the third molecule, the sulfate ion is triply coordinated to the metal center with one oxygen coordinated to both Mn(II) ions, one oxygen coordinated to Mn1, and one oxygen coordinated to Mn2. The sulfate in this coordination mode displaces the bridging ligand and one of the terminal solvent ligands. In both sulfate coordination modes, the sulfate ion is stabilized by hydrogen bonding interactions with conserved arginine residues, Arg 53 and Arg 162. The two different active site structures provide models for intermediates in phosphoester hydrolysis and suggest specific mechanistic roles for conserved residues.
- Department of Biochemistry, Northwestern University, Evanston, Illinois 60208, USA.
Organizational Affiliation: