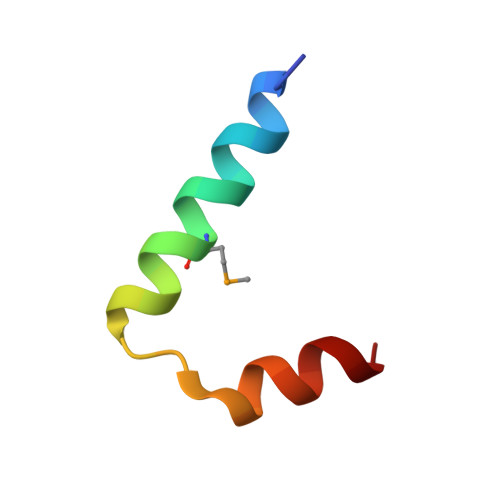
High-resolution structure of the HNF-1alpha dimerization domain.
Rose, R.B., Endrizzi, J.A., Cronk, J.D., Holton, J., Alber, T.(2000) Biochemistry 39: 15062-15070
- PubMed: 11106484 Search on PubMed
- DOI: https://doi.org/10.1021/bi001996t
- Primary Citation Related Structures:
1G2Y, 1G2Z, 1G39 - PubMed Abstract:
The N-terminal dimerization domain of the transcriptional activator hepatocyte nuclear factor-1alpha (HNF-1alpha) is essential for DNA binding and association of the transcriptional coactivator, DCoH (dimerization cofactor of HNF-1). To investigate the basis for dimerization of HNF-1 proteins, we determined the 1.2 A resolution X-ray crystal structure of the dimerization domain of HNF-1alpha (HNF-p1). Phasing was facilitated by devising a simple synthesis for Fmoc-selenomethionine and substituting leucine residues with selenomethionine. The HNF-1 dimerization domain forms a unique, four-helix bundle that is preserved with localized conformational shifts in the DCoH complex. In three different crystal forms, HNF-p1 displays subtle shifts in the conformation of the interhelix loop and the crossing angle between the amino- and carboxyl-terminal helices. In all three crystal forms, the HNF-p1 dimers pair through an exposed hydrophobic surface that also forms the binding site for DCoH. Conserved core residues in the dimerization domain of the homologous transcriptional regulator HNF-1beta rationalize the functional heterodimerization of the HNF-1alpha and HNF-1beta proteins. Mutations in HNF-1alpha are associated with maturity-onset diabetes of the young type 3 (MODY3), and the structure of HNF-p1 provides insights into the effects of three MODY3 mutations.
- Department of Molecular and Cell Biology, 229 Stanley Hall #3206, University of California, Berkeley, California 94720-3206, USA.
Organizational Affiliation: