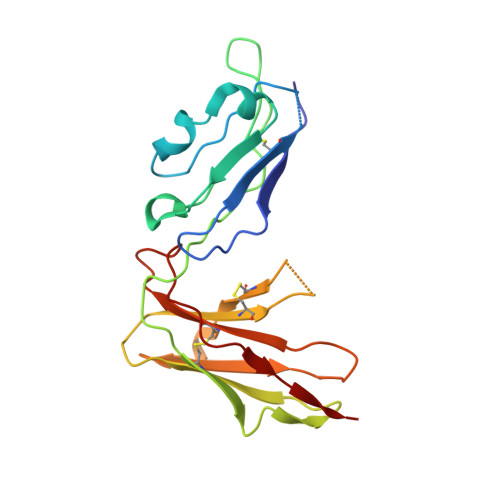
Crystal structure and ligand binding properties of the D1D2 region of the inhibitory receptor LIR-1 (ILT2).
Chapman, T.L., Heikema, A.P., West Jr., A.P., Bjorkman, P.J.(2000) Immunity 13: 727-736
- PubMed: 11114384 Search on PubMed
- DOI: https://doi.org/10.1016/s1074-7613(00)00071-6
- Primary Citation Related Structures:
1G0X - PubMed Abstract:
LIR-1 is an inhibitory receptor that recognizes class I MHC molecules and the human cytomegalovirus class I homolog UL18. Here, we report the 2.1 A resolution crystal structure of the ligand binding portion of LIR-1 (domains 1 and 2 [D1D2]) and localize the binding region for UL18. LIR-1 D1D2 is composed of two immunoglobulin-like domains arranged at an acute angle to form a bent structure resembling the structures of natural killer inhibitory receptors (KIRs). The LIR-1 binding site comprises a portion of D1 distant from the interdomain hinge region that constitutes the KIR binding site, consistent with differences in LIR-1 and KIR recognition properties and functions.
- Division of Biology 156-29, California Institute of Technology, Pasadena, CA 91125, USA.
Organizational Affiliation: