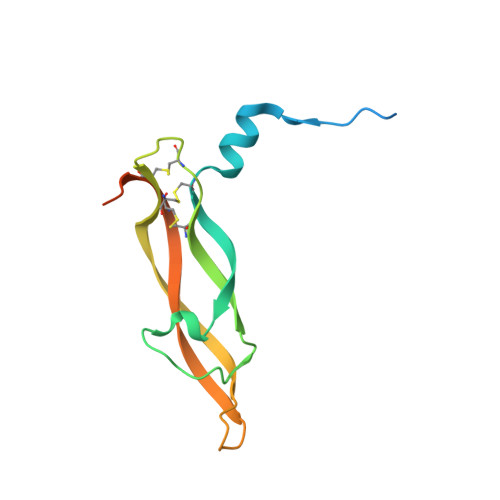
The crystal structure of human placenta growth factor-1 (PlGF-1), an angiogenic protein, at 2.0 A resolution.
Iyer, S., Leonidas, D.D., Swaminathan, G.J., Maglione, D., Battisti, M., Tucci, M., Persico, M.G., Acharya, K.R.(2001) J Biological Chem 276: 12153-12161
- PubMed: 11069911 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M008055200
- Primary Citation Related Structures:
1FZV - PubMed Abstract:
The angiogenic molecule placenta growth factor (PlGF) is a member of the cysteine-knot family of growth factors. In this study, a mature isoform of the human PlGF protein, PlGF-1, was crystallized as a homodimer in the crystallographic asymmetric unit, and its crystal structure was elucidated at 2.0 A resolution. The overall structure of PlGF-1 is similar to that of vascular endothelial growth factor (VEGF) with which it shares 42% amino acid sequence identity. Based on structural and biochemical data, we have mapped several important residues on the PlGF-1 molecule that are involved in recognition of the fms-like tyrosine kinase receptor (Flt-1, also known as VEGFR-1). We propose a model for the association of PlGF-1 and Flt-1 domain 2 with precise shape complementarity, consider the relevance of this assembly for PlGF-1 signal transduction, and provide a structural basis for altered specificity of this molecule.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, United Kingdom.
Organizational Affiliation: