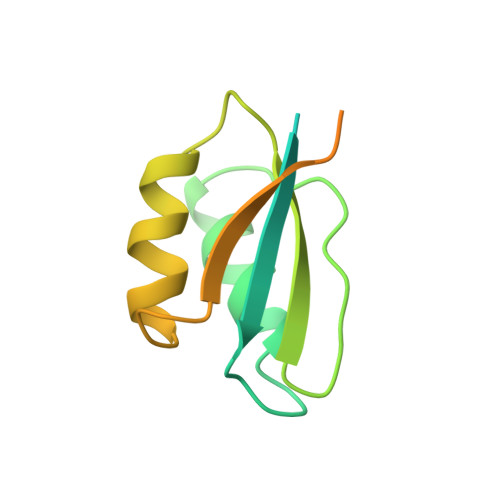
Structure and dynamics of a CheY-binding domain of the chemotaxis kinase CheA determined by nuclear magnetic resonance spectroscopy.
McEvoy, M.M., Muhandiram, D.R., Kay, L.E., Dahlquist, F.W.(1996) Biochemistry 35: 5633-5640
- PubMed: 8639521 Search on PubMed
- DOI: https://doi.org/10.1021/bi952707h
- Primary Citation Related Structures:
1FWP - PubMed Abstract:
The Escherichia coli histidine autokinase CheA plays an important role in coupling signals received from membrane-bound receptors to changes in the swimming behavior of the cells in order to respond appropriately to environmental signals. Here we describe the structure of the 14 kDa fragment of the chemotaxis kinase CheA, residues 124--257, which binds to the downstream targets of phosphorylation, the response regulators CheY and CheB. This protein fragment contains the CheY-binding domain flanked on each side by regions that correspond to domain linkers in the intact protein. The structure of the domain was determined from 1429 restraints derived from heteronuclear multidimensional NMR experiments. Hybrid distance geometry--dynamical simulated annealing methods were used to calculate a family of structures that satisfy the experimental distance restraints and torsion angle restraints. The root mean square deviation of the 69 ordered residues in the domain is 0.52 A for the backbone heavy atoms and 0.99 A for all heavy atoms. The residues that have been implicated as important for CheY binding form a face consisting of several partially buried hydrophobic residues, framed by charged residues. The dynamic properties of this protein fragment were measured and analyzed using both isotropic and anisotropic models of molecular motion. The linker regions are very flexible and disordered, as evidenced by the very dynamics properties as compared to the CheY-binding domain. The CheY-binding domain of CheA is structurally similar to the histidine-containing phosphocarrier, HPr, which is a protein involved in the phosphoenolpyruvate:sugar phosphotransferase (PTS) pathway. This structural similarity suggests a possible evolutionary relationship of the PTS and chemotaxis pathways.
- Institute of Molecular Biology, University of Oregon, Eugene 97403, USA.
Organizational Affiliation: