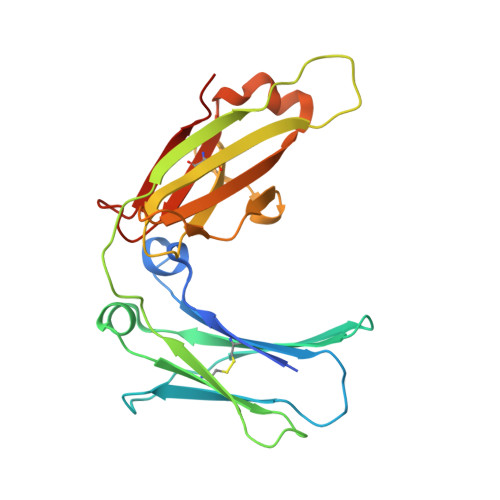
Structure of the human IgE-Fc C epsilon 3-C epsilon 4 reveals conformational flexibility in the antibody effector domains.
Wurzburg, B.A., Garman, S.C., Jardetzky, T.S.(2000) Immunity 13: 375-385
- PubMed: 11021535 Search on PubMed
- DOI: https://doi.org/10.1016/s1074-7613(00)00037-6
- Primary Citation Related Structures:
1FP5 - PubMed Abstract:
IgE antibodies mediate antiparasitic immune responses and the inflammatory reactions of allergy and asthma. We have solved the crystal structure of the human IgE-Fc Cepsilon3-Cepsilon4 domains to 2.3 A resolution. The structure reveals a large rearrangement of the N-terminal Cepsilon3 domains when compared to related IgG-Fc structures and to the IgE-Fc bound to its high-affinity receptor, FcepsilonRI. The IgE-Fc adopts a more compact, closed configuration that places the two Cepsilon3 domains in close proximity, decreases the size of the interdomain cavity, and obscures part of the FcepsilonRI binding site. IgE-Fc conformational flexibility may be required for interactions with two distinct IgE receptors, and the structure suggests strategies for the design of therapeutic compounds for the treatment of IgE-mediated diseases.
- Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University, Evanston, Illinois 60208, USA.
Organizational Affiliation: