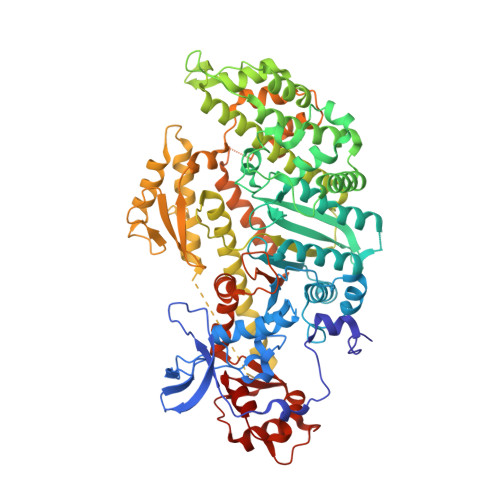
X-ray structures of the apo and MgATP-bound states of Dictyostelium discoideum myosin motor domain.
Bauer, C.B., Holden, H.M., Thoden, J.B., Smith, R., Rayment, I.(2000) J Biological Chem 275: 38494-38499
- PubMed: 10954715 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M005585200
- Primary Citation Related Structures:
1FMV, 1FMW - PubMed Abstract:
Myosin is the most comprehensively studied molecular motor that converts energy from the hydrolysis of MgATP into directed movement. Its motile cycle consists of a sequential series of interactions between myosin, actin, MgATP, and the products of hydrolysis, where the affinity of myosin for actin is modulated by the nature of the nucleotide bound in the active site. The first step in the contractile cycle occurs when ATP binds to actomyosin and releases myosin from the complex. We report here the structure of the motor domain of Dictyostelium discoideum myosin II both in its nucleotide-free state and complexed with MgATP. The structure with MgATP was obtained by soaking the crystals in substrate. These structures reveal that both the apo form and the MgATP complex are very similar to those previously seen with MgATPgammaS and MgAMP-PNP. Moreover, these structures are similar to that of chicken skeletal myosin subfragment-1. The crystallized protein is enzymatically active in solution, indicating that the conformation of myosin observed in chicken skeletal myosin subfragment-1 is unable to hydrolyze ATP and most likely represents the pre-hydrolysis structure for the myosin head that occurs after release from actin.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706-1544, USA.
Organizational Affiliation: