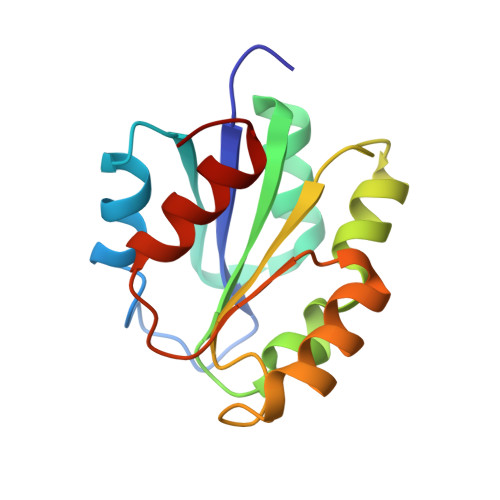
A protein pre-organized to trap the nucleotide moiety of coenzyme B(12): refined solution structure of the B(12)-binding subunit of glutamate mutase from Clostridium tetanomorphum.
Hoffmann, B., Tollinger, M., Konrat, R., Huhta, M., Marsh, E.N., Krautler, B.(2001) Chembiochem 2: 643-655
- PubMed: 11828501 Search on PubMed
- DOI: https://doi.org/10.1002/1439-7633(20010903)2:9<643::AID-CBIC643>3.0.CO;2-J
- Primary Citation Related Structures:
1FMF - PubMed Abstract:
Uniformly (13)C,(15)N-labeled MutS, the coenzyme B(12)-binding subunit of glutamate mutase from Clostridium tetanomorphum, was prepared by overexpression from an Escherichia coli strain. Multidimensional heteronuclear NMR spectroscopic experiments with aqueous solutions of (13)C,(15)N-labeled MutS provided signal assignments for roughly 90% of the 1025 hydrogen, 651 carbon, and 173 nitrogen atoms and resulted in about 1800 experimental restraints. Based on the information from the NMR experiments, the structure of MutS was calculated, confirming the earlier, less detailed structure obtained with (15)N-labeled MutS. The refined analysis allowed a precise determination of the secondary and tertiary structure including several crucial side chain interactions. The structures of (the apoprotein) MutS in solution and of the B(12)-binding subunit in the crystal of the corresponding homologous holoenzyme from Clostridium cochlearium differ only in a section that forms the well-structured helix alpha1 in the crystal structure and that also comprises the cobalt-coordinating histidine residue. In the apoprotein MutS, this part of the B(12)-binding subunit is dynamic. The carboxy-terminal end of this section is conformationally flexible and has significant propensity for an alpha-helical structure ("nascent helix"). This dynamic section in MutS is a decisive element for the binding of the nucleotide moiety of coenzyme B(12) and appears to be stabilized as a helix (alpha1) upon trapping of the nucleotide of the B(12) cofactor.
- Institute of Organic Chemistry, University of Innsbruck, Innrain 52a, 6020 Innsbruck, Austria.
Organizational Affiliation: