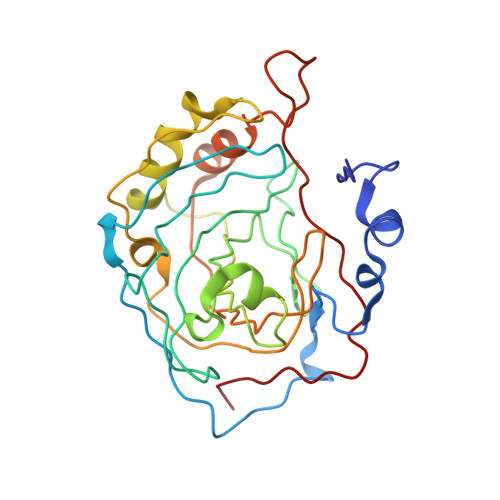
Crystal structure of S-glutathiolated carbonic anhydrase III.
Mallis, R.J., Poland, B.W., Chatterjee, T.K., Fisher, R.A., Darmawan, S., Honzatko, R.B., Thomas, J.A.(2000) FEBS Lett 482: 237-241
- PubMed: 11024467 Search on PubMed
- DOI: https://doi.org/10.1016/s0014-5793(00)02022-6
- Primary Citation Related Structures:
1FLJ - PubMed Abstract:
S-Glutathiolation of carbonic anhydrase III (CAIII) occurs rapidly in hepatocytes under oxidative stress. The crystal structure of the S-glutathiolated CAIII from rat liver reveals covalent adducts on cysteines 183 and 188. Electrostatic charge and steric contacts at each modification site inversely correlate with the relative rates of reactivity of these cysteines toward glutathione (GSH). Diffuse electron density associated with the GSH adducts suggests a lack of preferred bonding interactions between CAIII and the glutathionyl moieties. Hence, the GSH adducts are available for binding by a protein capable of reducing this mixed disulfide. These properties are consistent with the participation of CAIII in the protection/recovery from the damaging effects of oxidative agents.
- Department of Biochemistry, Biophysics and Molecular Biology, Iowa State University, Ames, IA 50011, USA.
Organizational Affiliation: