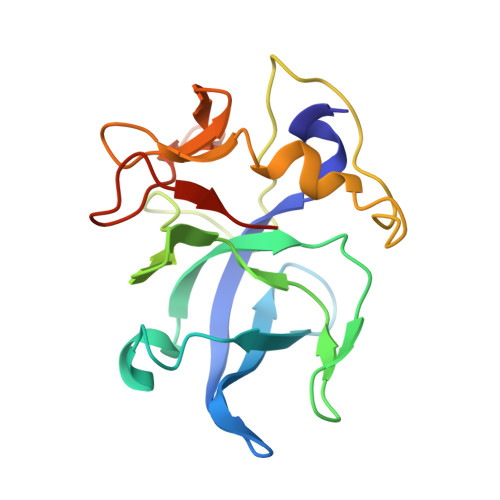
Structure of the EMAPII domain of human aminoacyl-tRNA synthetase complex reveals evolutionary dimer mimicry.
Renault, L., Kerjan, P., Pasqualato, S., Menetrey, J., Robinson, J.C., Kawaguchi, S., Vassylyev, D.G., Yokoyama, S., Mirande, M., Cherfils, J.(2001) EMBO J 20: 570-578
- PubMed: 11157763 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/20.3.570
- Primary Citation Related Structures:
1E7Z, 1FL0 - PubMed Abstract:
The EMAPII (endothelial monocyte-activating polypeptide II) domain is a tRNA-binding domain associated with several aminoacyl-tRNA synthetases, which becomes an independent domain with inflammatory cytokine activity upon apoptotic cleavage from the p43 component of the multisynthetase complex. It comprises a domain that is highly homologous to bacterial tRNA-binding proteins (Trbp), followed by an extra domain without homology to known proteins. Trbps, which may represent ancient tRNA chaperones, form dimers and bind one tRNA per dimer. In contrast, EMAPII domains are monomers. Here we report the crystal structure at 1.14 Angstroms of human EMAPII. The structure reveals that the Trbp-like domain, which forms an oligonucleotide-binding (OB) fold, is related by degenerate 2-fold symmetry to the extra-domain. The pseudo-axis coincides with the dyad axis of bacterial TtCsaA, a Trbp whose structure was solved recently. The interdomain interface in EMAPII mimics the intersubunit interface in TtCsaA, and may thus generate a novel OB-fold-based tRNA-binding site. The low sequence homology between the extra domain of EMAPII and either its own OB fold or that of Trbps suggests that dimer mimicry originated from convergent evolution rather than gene duplication.
- Laboratoire d'Enzymologie et Biochimie Structurales, CNRS, 1 Avenue de la Terrasse, 91198 Gif sur Yvette cedex, France.
Organizational Affiliation: