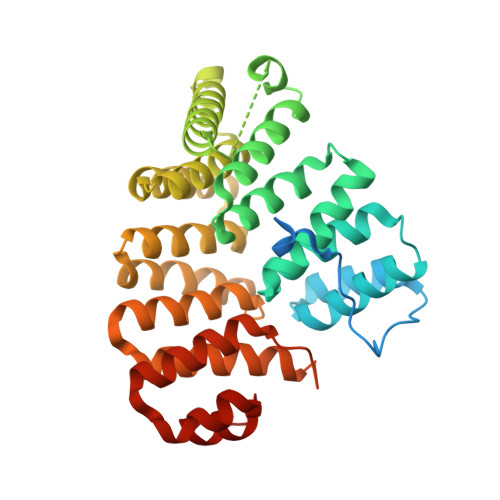
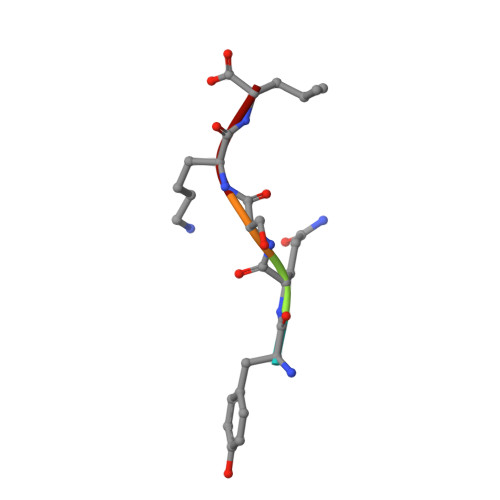
Peroxisomal targeting signal-1 recognition by the TPR domains of human PEX5.
Gatto Jr., G.J., Geisbrecht, B.V., Gould, S.J., Berg, J.M.(2000) Nat Struct Biol 7: 1091-1095
- PubMed: 11101887 Search on PubMed
- DOI: https://doi.org/10.1038/81930
- Primary Citation Related Structures:
1FCH - PubMed Abstract:
Many proteins contain targeting signals within their sequences that specify their delivery to particular organelles. The peroxisomal targeting signal-1 (PTS1) is a C-terminal tripeptide that is sufficient to direct proteins into peroxisomes. The PTS1 sequence closely approximates Ser-Lys-Leu-COO-. PEX5, the receptor for PTS1, interacts with the signal via a series of tetratricopeptide repeats (TPRs) within its C-terminal half. Here we report the crystal structure of a fragment of human PEX5 that includes all seven predicted TPR motifs in complex with a pentapeptide containing a PTS1 sequence. Two clusters of three TPRs almost completely surround the peptide, while a hinge region, previously identified as TPR4, forms a distinct structure that enables the two sets of TPRs to form a single binding site. This structure reveals the molecular basis for PTS1 recognition and demonstrates a novel mode of TPR-peptide interaction.
- Department of Biophysics and Biophysical Chemistry, The Johns Hopkins University School of Medicine, Baltimore, Maryland 21205, USA.
Organizational Affiliation: