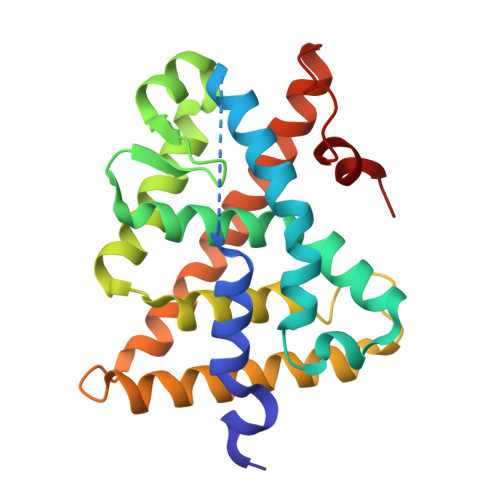
Crystal structure of the human RXRalpha ligand-binding domain bound to its natural ligand: 9-cis retinoic acid.
Egea, P.F., Mitschler, A., Rochel, N., Ruff, M., Chambon, P., Moras, D.(2000) EMBO J 19: 2592-2601
- PubMed: 10835357 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/19.11.2592
- Primary Citation Related Structures:
1FBY - PubMed Abstract:
The pleiotropic effects of active retinoids are transduced by their cognate nuclear receptors, retinoid X receptors (RXRs) and retinoic acid receptors (RARs), which act as transcriptional regulators activated by two stereoisomers of retinoic acid (RA): 9-cis RA (9-cRA) and all-trans RA (a-tRA). Among nuclear receptors, RXR occupies a central position and plays a crucial role in many intracellular signalling pathways as a ubiquitous heterodimerization partner with numerous other members of this superfamily. Whereas RARs bind both isomers, RXRs exclusively bind 9-cRA. The crystal structure of the ligand-binding domain (LBD) of human RXRalpha bound to 9-cRA reveals the molecular basis of this ligand selectivity and allows a comparison of both apo and holo forms of the same nuclear receptor. In the crystal, the receptor is monomeric and exhibits a canonical agonist conformation without direct contacts between the ligand and the transactivation helix H12. Comparison with the unliganded RXRalpha LBD structure reveals the molecular mechanisms of ligand-induced conformational changes and allows us to describe at the atomic level how these changes generate the proper protein interface involved in nuclear receptor-coactivator interaction.
- Institut de Génétique et de Biologie Moléculaire et Cellulaire, CNRS/INSERM/ULP/Collège de France, BP 163-67404 Illkirch Cedex, CU de Strasbourg, France.
Organizational Affiliation: