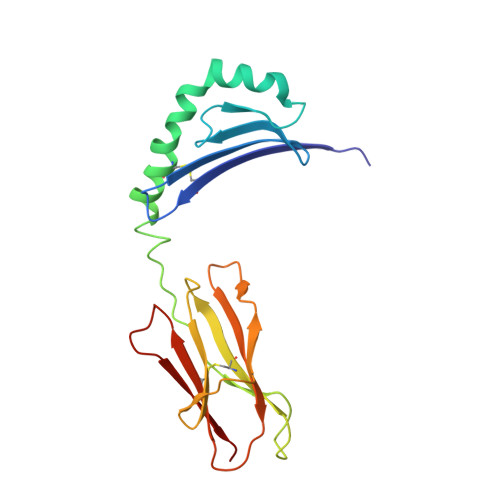
Structural basis of peptide binding and presentation by the type I diabetes-associated MHC class II molecule of NOD mice.
Latek, R.R., Suri, A., Petzold, S.J., Nelson, C.A., Kanagawa, O., Unanue, E.R., Fremont, D.H.(2000) Immunity 12: 699-710
- PubMed: 10894169 Search on PubMed
- DOI: https://doi.org/10.1016/s1074-7613(00)80220-4
- Primary Citation Related Structures:
1F3J - PubMed Abstract:
We have determined the crystal structure of I-Ag7, an integral component in murine type I diabetes development. Several features distinguish I-Ag7 from other non-autoimmune-associated MHC class II molecules, including novel peptide and heterodimer pairing interactions. The binding groove of I-Ag7 is unusual at both terminal ends, with a potentially solvent-exposed channel at the base of the P1 pocket and a widened entrance to the P9 pocket. Peptide binding studies with variants of the hen egg lysozyme I-Ag7 epitope HEL(11-25) support a comprehensive structure-based I-Ag7 binding motif. Residues critical for T cell recognition were investigated with a panel of HEL(11-25)-restricted clones, which uncovered P1 anchor-dependent structural variations. These results establish a framework for future experiments directed at understanding the role of I-Ag7 in autoimmunity.
- Department of Pathology and Immunology, Washington University School of Medicine, St. Louis, Missouri 63110, USA.
Organizational Affiliation: