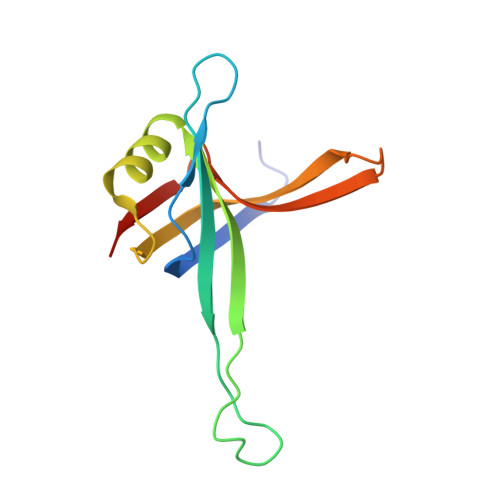
Structure of the DNA binding domain of E. coli SSB bound to ssDNA.
Raghunathan, S., Kozlov, A.G., Lohman, T.M., Waksman, G.(2000) Nat Struct Biol 7: 648-652
- PubMed: 10932248 Search on PubMed
- DOI: https://doi.org/10.1038/77943
- Primary Citation Related Structures:
1EYG - PubMed Abstract:
The structure of the homotetrameric DNA binding domain of the single stranded DNA binding protein from Escherichia coli (Eco SSB) bound to two 35-mer single stranded DNAs was determined to a resolution of 2.8 A. This structure describes the vast network of interactions that results in the extensive wrapping of single stranded DNA around the SSB tetramer and suggests a structural basis for its various binding modes.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, Campus Box 8231, 660 S. Euclid Ave., Saint Louis, Missouri 63110, USA.
Organizational Affiliation: