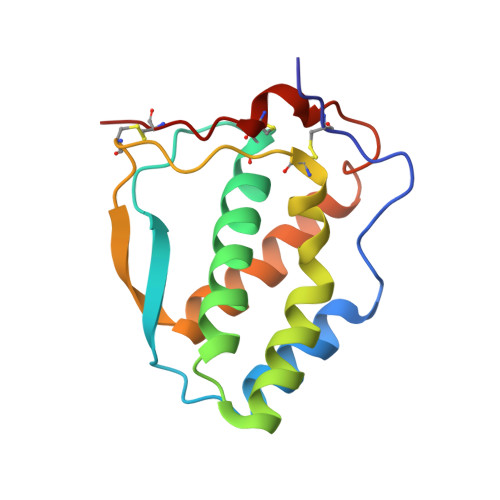
Flt3 ligand structure and unexpected commonalities of helical bundles and cystine knots.
Savvides, S.N., Boone, T., Andrew Karplus, P.(2000) Nat Struct Biol 7: 486-491
- PubMed: 10881197 Search on PubMed
- DOI: https://doi.org/10.1038/75896
- Primary Citation Related Structures:
1ETE - PubMed Abstract:
Human Flt3 ligand (Flt3L) stimulates early hematopoiesis by activating a type III tyrosine kinase receptor on primitive bone marrow stem cells. The crystal structure of soluble Flt3L reveals that it is a homodimer of two short chain alpha-helical bundles. Comparisons of structure-function relationships of Flt3L with the homologous hematopoietic cytokines macrophage colony stimulating factor (MCSF) and stem cell factor (SCF) suggest that they have a common receptor binding mode that is distinct from the paradigm derived from the complex of growth hormone with its receptor. Furthermore, we identify recognition features common to all helical and cystine-knot protein ligands that activate type III tyrosine kinase receptors, and the closely related type V tyrosine kinase receptors.
- Program in Biophysics, Cornell University, Ithaca, NY 14853, USA.
Organizational Affiliation: