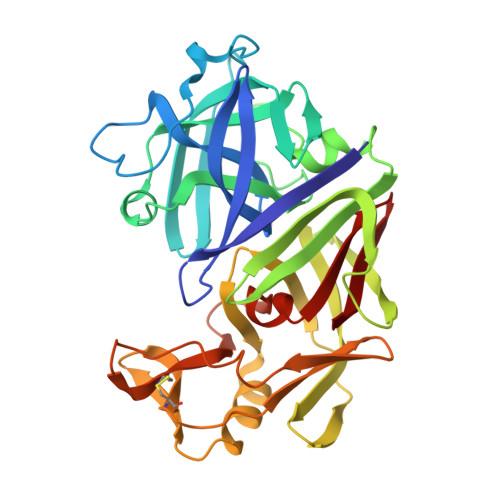
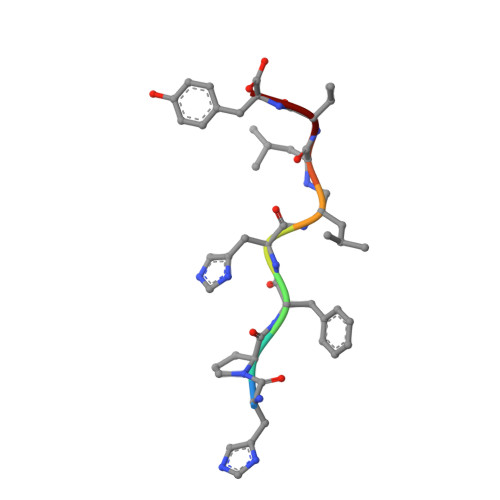
The Active Site of Aspartic Proteinases
Pearl, L., Blundell, T.(1984) FEBS Lett 174: 96-101
- PubMed: 6381096 Search on PubMed
- DOI: https://doi.org/10.1016/0014-5793(84)81085-6
- Primary Citation Related Structures:
1ER8, 3ER3, 4APE, 4ER1, 4ER2 - PubMed Abstract:
The active site of the aspartic proteinase, endothiapepsin, has been defined by X-ray analysis and restrained least-squares refinement at 2.1 A resolution with a crystallographic agreement value of 0.16. The environments of the two catalytically important aspartyl groups are remarkably similar and the contributions of the NH2- and COOH-terminal domains to the catalytic centre are related by a local 2-fold axis. The carboxylates of the aspartyls share a hydrogen bond and have equivalent contacts to a bound water molecule or hydroxonium ion lying on the local diad. The main chains around 32 and 215 are connected by a novel interaction involving diad-related threonines. It is suggested that the two pKa values of the active site aspartyls arise from a structure not unlike that in maleic acid with a hydrogen-bonded intermediate species and a dicarboxylate characterised by electrostatic repulsions between the two negatively charged groups.