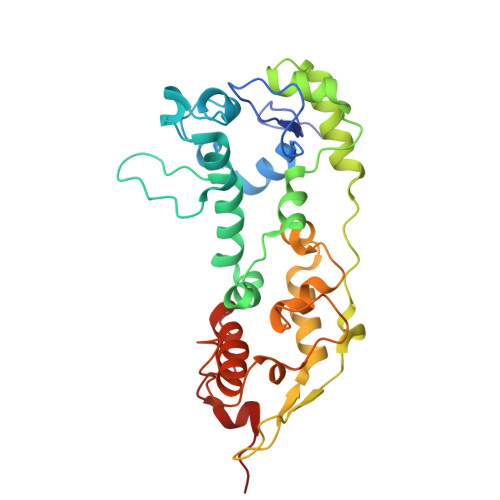
Three-dimensional structure of the nonaheme cytochrome c from Desulfovibrio desulfuricans Essex in the Fe(III) state at 1.89 A resolution.
Umhau, S., Fritz, G., Diederichs, K., Breed, J., Welte, W., Kroneck, P.M.(2001) Biochemistry 40: 1308-1316
- PubMed: 11170457 Search on PubMed
- DOI: https://doi.org/10.1021/bi001479a
- Primary Citation Related Structures:
1DUW - PubMed Abstract:
A nine heme group containing cytochrome c isolated from the soluble and membrane fractions of Desulfovibrio desulfuricans Essex, termed nonaheme cytochrome c, was crystallized, and the structure was solved using the multiple wavelength anomalous dispersion (MAD) phasing method. Refinement was carried out to a resolution of 1.89 A, and anisotropic temperature factors were addressed to the iron and sulfur atoms in the model. The structure revealed two cytochrome c(3) like domains with the typical arrangement of four heme centers. Both domains flanked an extra heme buried under the protein surface. This heme is held in position by loop extensions in each of the two domains. Although both the N- and C-terminal tetraheme domains exhibit a fold and heme arrangement very similar to that of cytochrome c(3), they differ considerably in their loop extensions and electrostatic surface. Analysis of the structure provides evidence for a different function of both domains, namely, anchoring the protein in a transmembranous complex with the N-terminal domain and formation of an electron-transfer complex with hydrogenase by the C-terminal domain.
- Fachbereich Biologie, Mathematisch-Naturwissenschaftliche Sektion, Universität Konstanz, 78457 Konstanz, Germany.
Organizational Affiliation: