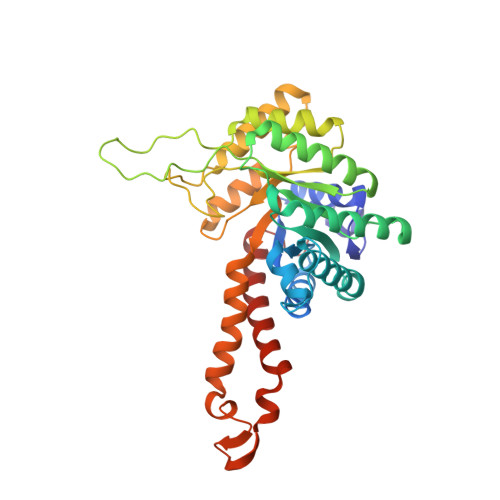
Novel active site in Escherichia coli fructose 1,6-bisphosphate aldolase.
Blom, N.S., Tetreault, S., Coulombe, R., Sygusch, J.(1996) Nat Struct Biol 3: 856-862
- PubMed: 8836102 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1096-856
- Primary Citation Related Structures:
1DOS - PubMed Abstract:
The molecular architecture of the Class II E. coli fructose 1,6-bisphosphate aldolase dimer was determined to 1.6 A resolution. The subunit fold corresponds to a singly wound alpha/beta-barrel with an active site located on the beta-barrel carboxyl side of each subunit. In each subunit there are two mutually exclusive zinc metal ion binding sites, 3.2 A apart; the exclusivity is mediated by a conformational transition involving side-chain rotations by chelating histidine residues. A binding site for K+ and NH4+ activators was found near the beta-barrel centre. Although Class I and Class II aldolases catalyse identical reactions, their active sites do not share common amino acid residues, are structurally dissimilar, and from sequence comparisons appear to be evolutionary distinct.
- Départment de biochimie, Université de Montréal, Canada.
Organizational Affiliation: