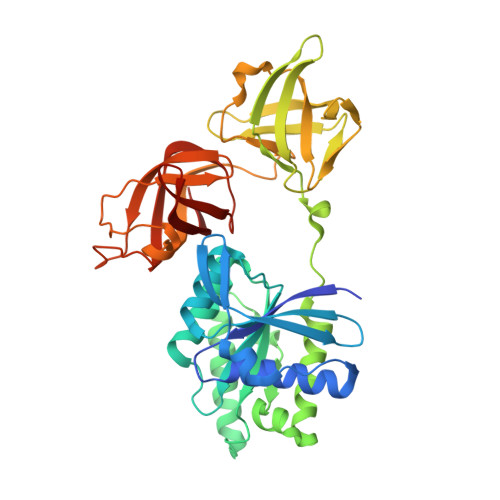
An alpha to beta conformational switch in EF-Tu.
Abel, K., Yoder, M.D., Hilgenfeld, R., Jurnak, F.(1996) Structure 4: 1153-1159
- PubMed: 8939740 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(96)00123-2
- Primary Citation Related Structures:
1DG1 - PubMed Abstract:
The bacterial elongation factor EF-Tu recognizes and transports aminoacyl-tRNAs to mRNA-programmed ribosomes. EF-Tu shares many structural and functional properties with other GTPases whose conformations are regulated by guanine nucleotides. An intact form of Escherichia coli EF-Tu complexed with GDP has been crystallized in the presence of the EF-Tu-specific antibiotic GE2270 A. The three-dimensional structure has been solved by X-ray diffraction analysis and refined to a final crystallographic R factor of 17.2% at a resolution of 2.5 A. The location of the GE2270 A antibiotic-binding site could not be identified. The structure of EF-Tu-GDP is nearly identical to that of a trypsin-modified form of EF-Tu-GDP, demonstrating conclusively that the protease treatment had not altered any essential structural features. The present structure represents the first view of an ordered Switch I region in EF-Tu-GDP and reveals similarities with two other GTPases complexed with GDP: Ran and ADP-ribosylation factor-1. A comparison of the Switch I regions of the GTP and GDP forms of EF-Tu also reveals that a segment, six amino acids in length, completely converts from an alpha helix in the GTP complex to beta secondary structure in the GDP form. The alpha to beta switch in EF-Tu may represent a prototypical activation mechanism for other protein families.
- Department of Biochemistry, University of California, Riverside, CA 92521-0129, USA.
Organizational Affiliation: