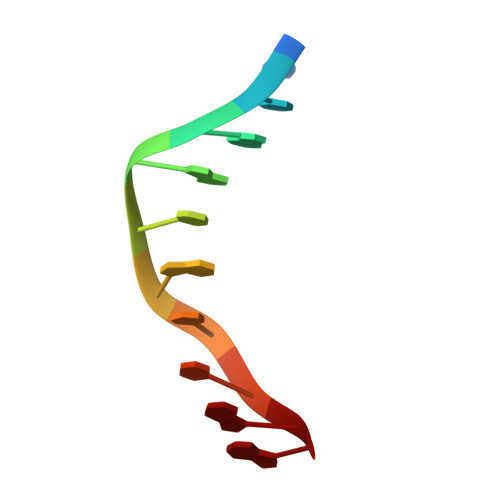
The Holliday junction in an inverted repeat DNA sequence: sequence effects on the structure of four-way junctions.
Eichman, B.F., Vargason, J.M., Mooers, B.H., Ho, P.S.(2000) Proc Natl Acad Sci U S A 97: 3971-3976
- PubMed: 10760268 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.97.8.3971
- Primary Citation Related Structures:
1DCV, 1DCW - PubMed Abstract:
Holliday junctions are important structural intermediates in recombination, viral integration, and DNA repair. We present here the single-crystal structure of the inverted repeat sequence d(CCGGTACCGG) as a Holliday junction at the nominal resolution of 2. 1 A. Unlike the previous crystal structures, this DNA junction has B-DNA arms with all standard Watson-Crick base pairs; it therefore represents the intermediate proposed by Holliday as being involved in homologous recombination. The junction is in the stacked-X conformation, with two interconnected duplexes formed by coaxially stacked arms, and is crossed at an angle of 41.4 degrees as a right-handed X. A sequence comparison with previous B-DNA and junction crystal structures shows that an ACC trinucleotide forms the core of a stable junction in this system. The 3'-C x G base pair of this ACC core forms direct and water-mediated hydrogen bonds to the phosphates at the crossover strands. Interactions within this core define the conformation of the Holliday junction, including the angle relating the stacked duplexes and how the base pairs are stacked in the stable form of the junction.
- Department of Biochemistry and Biophysics, ALS 2011, Oregon State University, Corvallis, OR 97331-7305, USA.
Organizational Affiliation: