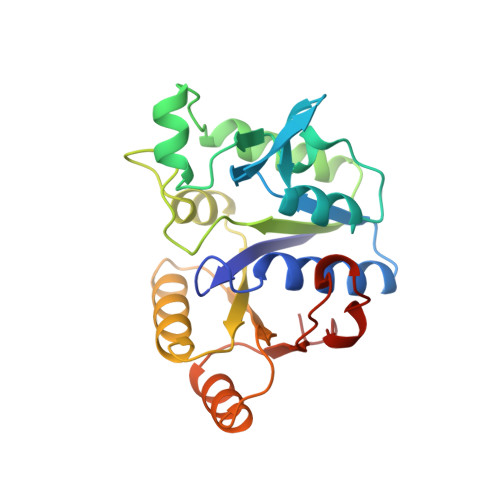
Snapshot of a phosphorylated substrate intermediate by kinetic crystallography.
Kack, H., Gibson, K.J., Lindqvist, Y., Schneider, G.(1998) Proc Natl Acad Sci U S A 95: 5495-5500
- PubMed: 9576910 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.95.10.5495
- Primary Citation Related Structures:
1A82, 1DAK - PubMed Abstract:
The ATP-dependent enzyme dethiobiotin synthetase from Escherichia coli catalyses the formation of dethiobiotin from CO2 and 7, 8-diaminopelargonic acid. The reaction is initiated by the formation of a carbamate and proceeds through a phosphorylated intermediate, a mixed carbamic phosphoric anhydride. Here, we report the crystal structures at 1.9- and 1.6-A resolution, respectively, of the enzyme-MgATP-diaminopelargonic acid and enzyme-MgADP-carbamic-phosphoric acid anhydride complexes, observed by using kinetic crystallography. Reaction initiation by addition of either NaHCO3 or diaminopelargonic acid to crystals already containing cosubstrates resulted in the accumulation of the phosphorylated intermediate at the active site. The phosphoryl transfer step shows inversion of the configuration at the phosphorus atom, consistent with an in-line attack by the carbamate oxygen onto the phosphorus atom of ATP. A key feature in the structure of the complex of the enzyme with the reaction intermediate is two magnesium ions, bridging the phosphates at the cleavage site. These magnesium ions compensate the negative charges at both phosphate groups after phosphoryl transfer and contribute to the stabilization of the reaction intermediate.
- Molecular Structural Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, 171 77 Stockholm, Sweden.
Organizational Affiliation: