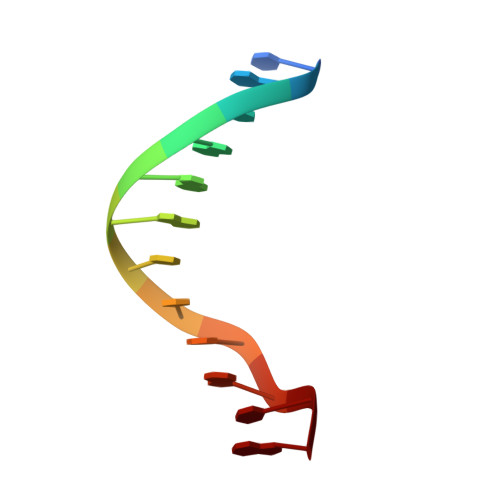
Crystal and molecular structure of a DNA fragment: d(CGTGAATTCACG).
Narayana, N., Ginell, S.L., Russu, I.M., Berman, H.M.(1991) Biochemistry 30: 4449-4455
- PubMed: 2021634 Search on PubMed
- DOI: https://doi.org/10.1021/bi00232a011
- Primary Citation Related Structures:
1D28 - PubMed Abstract:
The crystal structure of the dodecanucleotide d(CGTGAATTCACG) has been determined to a resolution of 2.7 A and refined to an R factor of 17.0% for 1532 reflections. The sequence crystallizes as a B-form double helix, with Watson-Crick base pairing. This sequence contains the EcoRI restriction endonuclease recognition site, GAATTC, and is flanked by CGT on the 5'-end and ACG on the 3'-end, in contrast to the CGC on the 5'-end and GCG on the 3'-end in the parent dodecamer d(CGCGAATTCGCG). A comparison with the isomorphous parent compound shows that any changes in the structure induced by the change in the sequence in the flanking region are highly localized. The global conformation of the duplex is conserved. The overall bend in the helix is 10 degrees. The average helical twist values for the present and the parent structures are 36.5 degrees and 36.4 degrees, respectively, corresponding to 10 base pairs per turn. The buckle at the substituted sites are significantly different from those seen at the corresponding positions in the parent dodecamer. Step 2 (GpT) is underwound with respect to the parent structure (27 degrees vs 36 degrees) and step 3 (TpG) is overwound (34 degrees vs 27 degrees). There is a spine of hydration in the narrow minor groove. The N3 atom of adenine on the substituted A10 and A22 bases are involved in the formation of hydrogen bonds with other duplexes or with water; the N3 atom of guanine on G10 and G22 bases in the parent structure does not form hydrogen bonds.
- Department of Chemistry, Rutgers University, Piscataway, New Jersey 08855.
Organizational Affiliation: