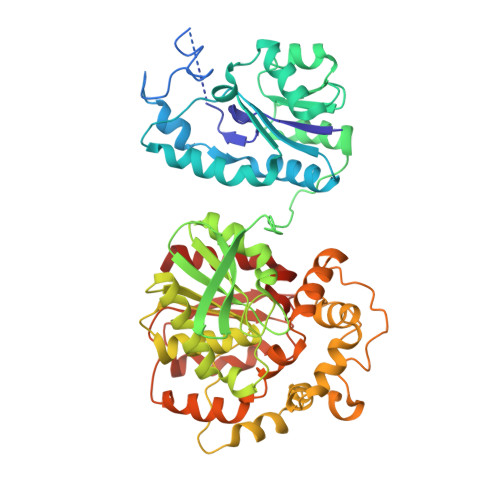
Detoxification of environmental mutagens and carcinogens: structure, mechanism, and evolution of liver epoxide hydrolase.
Argiriadi, M.A., Morisseau, C., Hammock, B.D., Christianson, D.W.(1999) Proc Natl Acad Sci U S A 96: 10637-10642
- PubMed: 10485878 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.96.19.10637
- Primary Citation Related Structures:
1CQZ, 1CR6 - PubMed Abstract:
The crystal structure of recombinant murine liver cytosolic epoxide hydrolase (EC 3.3.2.3) has been determined at 2.8-A resolution. The binding of a nanomolar affinity inhibitor confirms the active site location in the C-terminal domain; this domain is similar to that of haloalkane dehalogenase and shares the alpha/beta hydrolase fold. A structure-based mechanism is proposed that illuminates the unique chemical strategy for the activation of endogenous and man-made epoxide substrates for hydrolysis and detoxification. Surprisingly, a vestigial active site is found in the N-terminal domain similar to that of another enzyme of halocarbon metabolism, haloacid dehalogenase. Although the vestigial active site does not participate in epoxide hydrolysis, the vestigial domain plays a critical structural role by stabilizing the dimer in a distinctive domain-swapped architecture. Given the genetic and structural relationships among these enzymes of xenobiotic metabolism, a structure-based evolutionary sequence is postulated.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, PA 19104-6323, USA.
Organizational Affiliation: